Influenza virus hemagglutinin and neuraminidase, but not the matrix protein, are required for assembly and budding of plasmid-derived virus-like particles
- PMID: 17475660
- PMCID: PMC1933269
- DOI: 10.1128/JVI.00361-07
Influenza virus hemagglutinin and neuraminidase, but not the matrix protein, are required for assembly and budding of plasmid-derived virus-like particles
Abstract
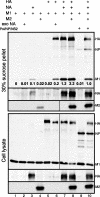
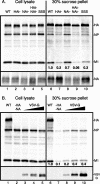
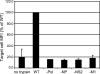
For influenza virus, we developed an efficient, noncytotoxic, plasmid-based virus-like particle (VLP) system to reflect authentic virus particles. This system was characterized biochemically by analysis of VLP protein composition, morphologically by electron microscopy, and functionally with a VLP infectivity assay. The VLP system was used to address the identity of the minimal set of viral proteins required for budding. Combinations of viral proteins were expressed in cells, and the polypeptide composition of the particles released into the culture media was analyzed. Contrary to previous findings in which matrix (M1) protein was considered to be the driving force of budding because M1 was found to be released copiously into the culture medium when M1 was expressed by using the vaccinia virus T7 RNA polymerase-driven overexpression system, in our noncytotoxic VLP system M1 was not released efficiently into the culture medium. Additionally, hemagglutinin (HA), when treated with exogenous neuraminidase (NA) or coexpressed with viral NA, could be released from cells independently of M1. Incorporation of M1 into VLPs required HA expression, although when M1 was omitted from VLPs, particles with morphologies similar to those of wild-type VLPs or viruses were observed. Furthermore, when HA and NA cytoplasmic tail mutants were included in the VLPs, M1 failed to be efficiently incorporated into VLPs, consistent with a model in which the glycoproteins control virus budding by sorting to lipid raft microdomains and recruiting the internal viral core components. VLP formation also occurred independently of the function of Vps4 in the multivesicular body pathway, as dominant-negative Vps4 proteins failed to inhibit influenza VLP budding.
Figures






References
-
- Barman, S., A. Ali, E. K.-W. Hui, L. Adhikary, and D. P. Nayak. 2001. Transport of viral proteins to the apical membranes and interaction of matrix proteins with glycoproteins in the assembly of influenza viruses. Virus Res. 77:61-69. - PubMed
-
- Bieniasz, P. D. 2006. Late budding domains and host proteins in enveloped virus release. Virology 344:55-63. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

