A Markov chain Monte Carlo approach for joint inference of population structure and inbreeding rates from multilocus genotype data
- PMID: 17483417
- PMCID: PMC1931536
- DOI: 10.1534/genetics.107.072371
A Markov chain Monte Carlo approach for joint inference of population structure and inbreeding rates from multilocus genotype data
Abstract
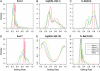
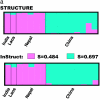
Nonrandom mating induces correlations in allelic states within and among loci that can be exploited to understand the genetic structure of natural populations (Wright 1965). For many species, it is of considerable interest to quantify the contribution of two forms of nonrandom mating to patterns of standing genetic variation: inbreeding (mating among relatives) and population substructure (limited dispersal of gametes). Here, we extend the popular Bayesian clustering approach STRUCTURE (Pritchard et al. 2000) for simultaneous inference of inbreeding or selfing rates and population-of-origin classification using multilocus genetic markers. This is accomplished by eliminating the assumption of Hardy-Weinberg equilibrium within clusters and, instead, calculating expected genotype frequencies on the basis of inbreeding or selfing rates. We demonstrate the need for such an extension by showing that selfing leads to spurious signals of population substructure using the standard STRUCTURE algorithm with a bias toward spurious signals of admixture. We gauge the performance of our method using extensive coalescent simulations and demonstrate that our approach can correct for this bias. We also apply our approach to understanding the population structure of the wild relative of domesticated rice, Oryza rufipogon, an important partially selfing grass species. Using a sample of n = 16 individuals sequenced at 111 random loci, we find strong evidence for existence of two subpopulations, which correlates well with geographic location of sampling, and estimate selfing rates for both groups that are consistent with estimates from experimental data (s approximately 0.48-0.70).
Figures





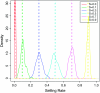
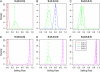
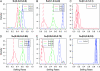
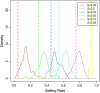

Similar articles
-
Estimating selfing rates from reconstructed pedigrees using multilocus genotype data.Mol Ecol. 2012 Jan;21(1):100-16. doi: 10.1111/j.1365-294X.2011.05373.x. Epub 2011 Nov 22. Mol Ecol. 2012. PMID: 22106925
-
A Markov chain Monte Carlo strategy for sampling from the joint posterior distribution of pedigrees and population parameters under a Fisher-Wright model with partial selfing.Theor Popul Biol. 2007 Nov;72(3):436-58. doi: 10.1016/j.tpb.2007.03.002. Epub 2007 Mar 12. Theor Popul Biol. 2007. PMID: 17448511 Review.
-
Maximum-likelihood and markov chain monte carlo approaches to estimate inbreeding and effective size from allele frequency changes.Genetics. 2003 Jul;164(3):1189-204. doi: 10.1093/genetics/164.3.1189. Genetics. 2003. PMID: 12871924 Free PMC article.
-
Performing Parentage Analysis in the Presence of Inbreeding and Null Alleles.Genetics. 2018 Dec;210(4):1467-1481. doi: 10.1534/genetics.118.301592. Epub 2018 Oct 18. Genetics. 2018. PMID: 30337340 Free PMC article.
-
Quantifying inbreeding in natural populations of hermaphroditic organisms.Heredity (Edinb). 2008 Apr;100(4):431-9. doi: 10.1038/hdy.2008.2. Epub 2008 Feb 27. Heredity (Edinb). 2008. PMID: 18301439 Review.
Cited by
-
Escape to Ferality: The Endoferal Origin of Weedy Rice from Crop Rice through De-Domestication.PLoS One. 2016 Sep 23;11(9):e0162676. doi: 10.1371/journal.pone.0162676. eCollection 2016. PLoS One. 2016. PMID: 27661982 Free PMC article.
-
Genetic diversity, seed size associations and population structure of a core collection of common beans (Phaseolus vulgaris L.).Theor Appl Genet. 2009 Oct;119(6):955-72. doi: 10.1007/s00122-009-1064-8. Epub 2009 Aug 18. Theor Appl Genet. 2009. PMID: 19688198
-
Fast and accurate population admixture inference from genotype data from a few microsatellites to millions of SNPs.Heredity (Edinb). 2022 Aug;129(2):79-92. doi: 10.1038/s41437-022-00535-z. Epub 2022 May 4. Heredity (Edinb). 2022. PMID: 35508539 Free PMC article.
-
Molecular Genetic Analysis with Microsatellite-like Loci Reveals Specific Dairy-Associated and Environmental Populations of the Yeast Geotrichum candidum.Microorganisms. 2022 Jan 4;10(1):103. doi: 10.3390/microorganisms10010103. Microorganisms. 2022. PMID: 35056553 Free PMC article.
-
Out to sea: ocean currents and patterns of asymmetric gene flow in an intertidal fish species.Front Genet. 2023 Jun 28;14:1206543. doi: 10.3389/fgene.2023.1206543. eCollection 2023. Front Genet. 2023. PMID: 37456662 Free PMC article.
References
-
- Ayres, K. L., and D. J. Balding, 1998. Measuring departures from Hardy-Weinberg: a Markov chain Monte Carlo method for estimating the inbreeding coefficient. Heredity 80(6): 769–777. - PubMed
-
- Dawson, K. J., and K. Belkhir, 2001. A Bayesian approach to the identification of panmictic populations and the assignment of individuals. Genet. Res. 78: 59–77. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical

