Quantitative gene expression assessment identifies appropriate cell line models for individual cervical cancer pathways
- PMID: 17493265
- PMCID: PMC1878486
- DOI: 10.1186/1471-2164-8-117
Quantitative gene expression assessment identifies appropriate cell line models for individual cervical cancer pathways
Abstract
Background: Cell lines have been used to study cancer for decades, but truly quantitative assessment of their performance as models is often lacking. We used gene expression profiling to quantitatively assess the gene expression of nine cell line models of cervical cancer.
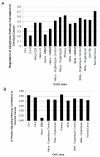
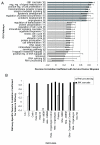
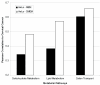
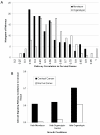
Results: We find a wide variation in the extent to which different cell culture models mimic late-stage invasive cervical cancer biopsies. The lowest agreement was from monolayer HeLa cells, a common cervical cancer model; the highest agreement was from primary epithelial cells, C4-I, and C4-II cell lines. In addition, HeLa and SiHa cell lines cultured in an organotypic environment increased their correlation to cervical cancer significantly. We also find wide variation in agreement when we considered how well individual biological pathways model cervical cancer. Cell lines with an anti-correlation to cervical cancer were also identified and should be avoided.
Conclusion: Using gene expression profiling and quantitative analysis, we have characterized nine cell lines with respect to how well they serve as models of cervical cancer. Applying this method to individual pathways, we identified the appropriateness of particular cell lines for studying specific pathways in cervical cancer. This study will allow researchers to choose a cell line with the highest correlation to cervical cancer at a pathway level. This method is applicable to other cancers and could be used to identify the appropriate cell line and growth condition to employ when studying other cancers.
Figures

References
-
- Goodwin EC, DiMaio D. Repression of human papillomavirus oncogenes in HeLa cervical carcinoma cells causes the orderly reactivation of dormant tumor suppressor pathways. Proceedings of the National Academy of Sciences of the United States of America. 2000;97:12513–12518. doi: 10.1073/pnas.97.23.12513. - DOI - PMC - PubMed
-
- Harima Y, Togashi A, Horikoshi K, Imamura M, Sougawa M, Sawada S, Tsunoda T, Nakamura Y, Katagiri T. Prediction of outcome of advanced cervical cancer to thermoradiotherapy according to expression profiles of 35 genes selected by cDNA microarray analysis. Int J Radiat Oncol Biol Phys. 2004;60:237–248. doi: 10.1016/j.ijrobp.2004.02.047. - DOI - PubMed
-
- Ahn WS, Huh SW, Bae SM, Lee IP, Lee JM, Namkoong SE, Kim CK, Sin JI. A major constituent of green tea, EGCG, inhibits the growth of a human cervical cancer cell line, CaSki cells, through apoptosis, G(1) arrest, and regulation of gene expression. DNA & Cell Biology. 2003;22:217–224. doi: 10.1089/104454903321655846. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

