A molecular dynamics study of slow base flipping in DNA using conformational flooding
- PMID: 17496048
- PMCID: PMC1913169
- DOI: 10.1529/biophysj.106.091751
A molecular dynamics study of slow base flipping in DNA using conformational flooding
Abstract
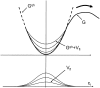
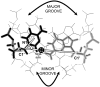
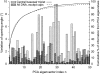
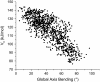
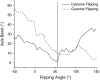
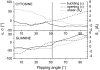
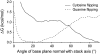
Individual DNA bases are known to be able to flip out of the helical stack, providing enzymes with access to the genetic information otherwise hidden inside the helix. Consequently, base flipping is a necessary first step to many more complex biological processes such as DNA transcription or replication. Much remains unknown about this elementary step, despite a wealth of experimental and theoretical studies. From the theoretical point of view, the involved timescale of milliseconds or longer requires the use of enhanced sampling techniques. In contrast to previous theoretical studies employing umbrella sampling along a predefined flipping coordinate, this study attempts to induce flipping without prior knowledge of the pathway, using information from a molecular dynamics simulation of a B-DNA fragment and the conformational flooding method. The relevance to base flipping of the principal components of the simulation is assayed, and a combination of modes optimally related to the flipping of the base through either helical groove is derived for each of the two bases of the central guanine-cytosine basepair. By applying an artificial flooding potential along these collective coordinates, the flipping mechanism is accelerated to within the scope of molecular dynamics simulations. The associated free energy surface is found to feature local minima corresponding to partially flipped states, particularly relevant to flipping in isolated DNA; further transitions from these minima to the fully flipped conformation are accelerated by additional flooding potentials. The associated free energy profiles feature similar barrier heights for both bases and pathways; the flipped state beyond is a broad and rugged attraction basin, only a few kcal/mol higher in energy than the closed conformation. This result diverges from previous works but echoes some aspects of recent experimental findings, justifying the need for novel approaches to this difficult problem: this contribution represents a first step in this direction. Important structural factors involved in flipping, both local (sugar-phosphate backbone dihedral angles) and global (helical axis bend), are also identified.
Figures


References
-
- Daujotyte, D., S. Serva, G. Vilkaitis, E. Merkiene, D. Venclovas, and S. Klimasauskas. 2004. Hhal DNA methyltransferase uses the protruding Gln237 for active flipping of its target cytosine. Structure. 12:1047–1055. - PubMed
-
- Klimasauskas, S., S. Kumar, R. J. Roberts, and X. D. Cheng. 1994. Hhal methyltransferase flips its target base out of the DNA helix. Cell. 76:357–369. - PubMed
-
- Klimasauskas, S., and R. J. Roberts. 1995. Disruption of the target G-C base-pair by the HhaI methyltransferase. Gene. 157:163–164. - PubMed
-
- Slupphaug, G., C. D. Mol, B. Kavli, A. S. Arvai, H. E. Krokan, and J. A. Tainer. 1996. A nucleotide-flipping mechanism from the structure of human uracil-DNA glycosylase bound to DNA. Nature. 384:87–92. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

