NrdR controls differential expression of the Escherichia coli ribonucleotide reductase genes
- PMID: 17496099
- PMCID: PMC1951866
- DOI: 10.1128/JB.00440-07
NrdR controls differential expression of the Escherichia coli ribonucleotide reductase genes
Abstract
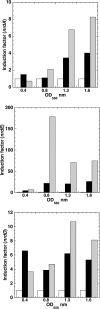
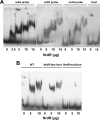
Escherichia coli possesses class Ia, class Ib, and class III ribonucleotide reductases (RNR). Under standard laboratory conditions, the aerobic class Ia nrdAB RNR genes are well expressed, whereas the aerobic class Ib nrdEF RNR genes are poorly expressed. The class III RNR is normally expressed under microaerophilic and anaerobic conditions. In this paper, we show that the E. coli YbaD protein differentially regulates the expression of the three sets of genes. YbaD is a homolog of the Streptomyces NrdR protein. It is not essential for growth and has been renamed NrdR. Previously, Streptomyces NrdR was shown to transcriptionally regulate RNR genes by binding to specific 16-bp sequence motifs, NrdR boxes, located in the regulatory regions of its RNR operons. All three E. coli RNR operons contain two such NrdR box motifs positioned in their regulatory regions. The NrdR boxes are located near to or overlap with the promoter elements. DNA binding experiments showed that NrdR binds to each of the upstream regulatory regions. We constructed deletions in nrdR (ybaD) and showed that they caused high-level induction of transcription of the class Ib RNR genes but had a much smaller effect on induction of transcription of the class Ia and class III RNR genes. We propose a model for differential regulation of the RNR genes based on binding of NrdR to the regulatory regions. The model assumes that differences in the positions of the NrdR binding sites, and in the sequences of the motifs themselves, determine the extent to which NrdR represses the transcription of each RNR operon.
Figures


References
-
- Aravind, L., Y. I. Wolf, and E. V. Koonin. 2000. The ATP-cone: an evolutionarily mobile, ATP-binding regulatory domain. J. Mol. Microbiol. Biotechnol. 2:191-194. - PubMed
-
- Atkin, C. L., L. Thelander, P. Reichard, and G. Lang. 1973. Iron and free radical in ribonucleotide reductase. Exchange of iron and Mossbauer spectroscopy of the protein B2 subunit of the Escherichia coli enzyme. J. Biol. Chem. 248:7464-7472. - PubMed
-
- Blattner, F. R., G. Plunkett III, C. A. Bloch, N. T. Perna, V. Burland, M. Riley, J. Collado-Vides, J. D. Glasner, C. K. Rode, G. F. Mayhew, J. Gregor, N. W. Davis, H. A. Kirkpatrick, M. A. Goeden, D. J. Rose, B. Mau, and Y. Shao. 1997. The complete genome sequence of Escherichia coli K-12. Science 277:1453-1474. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

