Dissociation of eIF1 from the 40S ribosomal subunit is a key step in start codon selection in vivo
- PMID: 17504939
- PMCID: PMC1865493
- DOI: 10.1101/gad.1528307
Dissociation of eIF1 from the 40S ribosomal subunit is a key step in start codon selection in vivo
Abstract
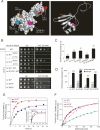
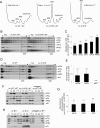
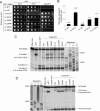
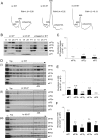
Selection of the AUG start codon is a key step in translation initiation requiring hydrolysis of GTP in the eIF2*GTP*Met-tRNA(i)(Met) ternary complex (TC) and subsequent P(i) release from eIF2*GDP*P(i). It is thought that eIF1 prevents recognition of non-AUGs by promoting scanning and blocking P(i) release at non-AUG codons. We show that Sui(-) mutations in Saccharomyces cerevisiae eIF1, which increase initiation at UUG codons, reduce interaction of eIF1 with 40S subunits in vitro and in vivo, and both defects are diminished in cells by overexpressing the mutant proteins. Remarkably, Sui(-) mutation ISQLG(93-97)ASQAA (abbreviated 93-97) accelerates eIF1 dissociation and P(i) release from reconstituted preinitiation complexes (PICs), whereas a hyperaccuracy mutation in eIF1A (that suppresses Sui(-) mutations) decreases the eIF1 off-rate. These findings demonstrate that eIF1 dissociation is a critical step in start codon selection, which is modulated by eIF1A. We also describe Gcd(-) mutations in eIF1 that impair TC loading on 40S subunits or destabilize the multifactor complex containing eIF1, eIF3, eIF5, and TC, showing that eIF1 promotes PIC assembly in vivo beyond its important functions in AUG selection.
Figures
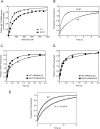
Comment in
-
Translation factor control of ribosome conformation during start codon selection.Genes Dev. 2007 Jun 1;21(11):1280-7. doi: 10.1101/gad.1562707. Genes Dev. 2007. PMID: 17545463 No abstract available.
References
-
- Algire M.A., Maag D., Savio P., Acker M.G., Tarun S.Z., Jr., Sachs A.B., Asano K., Nielsen K.H., Olsen D.S., Phan L., Maag D., Savio P., Acker M.G., Tarun S.Z., Jr., Sachs A.B., Asano K., Nielsen K.H., Olsen D.S., Phan L., Savio P., Acker M.G., Tarun S.Z., Jr., Sachs A.B., Asano K., Nielsen K.H., Olsen D.S., Phan L., Acker M.G., Tarun S.Z., Jr., Sachs A.B., Asano K., Nielsen K.H., Olsen D.S., Phan L., Tarun S.Z., Jr., Sachs A.B., Asano K., Nielsen K.H., Olsen D.S., Phan L., Sachs A.B., Asano K., Nielsen K.H., Olsen D.S., Phan L., Asano K., Nielsen K.H., Olsen D.S., Phan L., Nielsen K.H., Olsen D.S., Phan L., Olsen D.S., Phan L., Phan L., et al. Development and characterization of a reconstituted yeast translation initiation system. RNA. 2002;8:382–397. - PMC - PubMed
-
- Algire M.A., Maag D., Lorsch J.R., Maag D., Lorsch J.R., Lorsch J.R. π release from eIF2, not GTP hydrolysis, is the step controlled by start-site selection during eukaryotic translation initiation. Mol. Cell. 2005;20:251–262. - PubMed
-
- Asano K., Phan L., Anderson J., Hinnebusch A.G., Phan L., Anderson J., Hinnebusch A.G., Anderson J., Hinnebusch A.G., Hinnebusch A.G. Complex formation by all five homologues of mammalian translation initiation factor 3 subunits from yeast Saccharomyces cerevisiae. J. Biol. Chem. 1998;273:18573–18585. - PubMed
-
- Asano K., Clayton J., Shalev A., Hinnebusch A.G., Clayton J., Shalev A., Hinnebusch A.G., Shalev A., Hinnebusch A.G., Hinnebusch A.G. A multifactor complex of eukaryotic initiation factors eIF1, eIF2, eIF3, eIF5, and initiator tRNAMet is an important translation initiation intermediate in vivo. Genes & Dev. 2000;14:2534–2546. - PMC - PubMed
-
- Asano K., Shalev A., Phan L., Nielsen K., Clayton J., Valášek L., Donahue T.F., Hinnebusch A.G., Shalev A., Phan L., Nielsen K., Clayton J., Valášek L., Donahue T.F., Hinnebusch A.G., Phan L., Nielsen K., Clayton J., Valášek L., Donahue T.F., Hinnebusch A.G., Nielsen K., Clayton J., Valášek L., Donahue T.F., Hinnebusch A.G., Clayton J., Valášek L., Donahue T.F., Hinnebusch A.G., Valášek L., Donahue T.F., Hinnebusch A.G., Donahue T.F., Hinnebusch A.G., Hinnebusch A.G. Multiple roles for the carboxyl terminal domain of eIF5 in translation initiation complex assembly and GTPase activation. EMBO J. 2001;20:2326–2337. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous
