PdhR (pyruvate dehydrogenase complex regulator) controls the respiratory electron transport system in Escherichia coli
- PMID: 17513468
- PMCID: PMC1951801
- DOI: 10.1128/JB.00229-07
PdhR (pyruvate dehydrogenase complex regulator) controls the respiratory electron transport system in Escherichia coli
Abstract
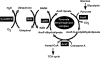
The pyruvate dehydrogenase (PDH) multienzyme complex plays a key role in the metabolic interconnection between glycolysis and the citric acid cycle. Transcription of the Escherichia coli genes for all three components of the PDH complex in the pdhR-aceEF-lpdA operon is repressed by the pyruvate-sensing PdhR, a GntR family transcription regulator, and derepressed by pyruvate. After a systematic search for the regulation targets of PdhR using genomic systematic evolution of ligands by exponential enrichment (SELEX), we have identified two novel targets, ndh, encoding NADH dehydrogenase II, and cyoABCDE, encoding the cytochrome bo-type oxidase, both together forming the pathway of respiratory electron transport downstream from the PDH cycle. PDH generates NADH, while Ndh and CyoABCDE together transport electrons from NADH to oxygen. Using gel shift and DNase I footprinting assays, the PdhR-binding site (PdhR box) was defined, which includes a palindromic consensus sequence, ATTGGTNNNACCAAT. The binding in vitro of PdhR to the PdhR box decreased in the presence of pyruvate. Promoter assays in vivo using a two-fluorescent-protein vector also indicated that the newly identified operons are repressed by PdhR and derepressed by the addition of pyruvate. Taken together, we propose that PdhR is a master regulator for controlling the formation of not only the PDH complex but also the respiratory electron transport system.
Figures






References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

