Gene cloning and characterization of the very large NAD-dependent l-glutamate dehydrogenase from the psychrophile Janthinobacterium lividum, isolated from cold soil
- PMID: 17526698
- PMCID: PMC1951823
- DOI: 10.1128/JB.00496-07
Gene cloning and characterization of the very large NAD-dependent l-glutamate dehydrogenase from the psychrophile Janthinobacterium lividum, isolated from cold soil
Abstract
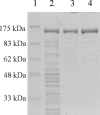
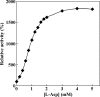
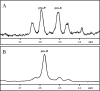
NAD-dependent l-glutamate dehydrogenase (NAD-GDH) activity was detected in cell extract from the psychrophile Janthinobacterium lividum UTB1302, which was isolated from cold soil and purified to homogeneity. The native enzyme (1,065 kDa, determined by gel filtration) is a homohexamer composed of 170-kDa subunits (determined by sodium dodecyl sulfate-polyacrylamide gel electrophoresis). Consistent with these findings, gene cloning and sequencing enabled deduction of the amino acid sequence of the subunit, which proved to be comprised of 1,575 amino acids with a combined molecular mass of 169,360 Da. The enzyme from this psychrophile thus appears to belong to the GDH family characterized by very large subunits, like those expressed by Streptomyces clavuligerus and Pseudomonas aeruginosa (about 180 kDa). The entire amino acid sequence of the J. lividum enzyme showed about 40% identity with the sequences from S. clavuligerus and P. aeruginosa enzymes, but the central domains showed higher homology (about 65%). Within the central domain, the residues related to substrate and NAD binding were highly conserved, suggesting that this is the enzyme's catalytic domain. In the presence of NAD, but not in the presence of NADP, this GDH exclusively catalyzed the oxidative deamination of l-glutamate. The stereospecificity of the hydride transfer to NAD was pro-S, which is the same as that of the other known GDHs. Surprisingly, NAD-GDH activity was markedly enhanced by the addition of various amino acids, such as l-aspartate (1,735%) and l-arginine (936%), which strongly suggests that the N- and/or C-terminal domains play regulatory roles and are involved in the activation of the enzyme by these amino acids.
Figures



Similar articles
-
The gdhB gene of Pseudomonas aeruginosa encodes an arginine-inducible NAD(+)-dependent glutamate dehydrogenase which is subject to allosteric regulation.J Bacteriol. 2001 Jan;183(2):490-9. doi: 10.1128/JB.183.2.490-499.2001. J Bacteriol. 2001. PMID: 11133942 Free PMC article.
-
Identification of catalytic residues of a very large NAD-glutamate dehydrogenase from Janthinobacterium lividum by site-directed mutagenesis.Biosci Biotechnol Biochem. 2014;78(12):2045-50. doi: 10.1080/09168451.2014.946394. Epub 2014 Aug 15. Biosci Biotechnol Biochem. 2014. PMID: 25126984
-
A new class of glutamate dehydrogenases (GDH). Biochemical and genetic characterization of the first member, the AMP-requiring NAD-specific GDH of Streptomyces clavuligerus.J Biol Chem. 2000 Dec 15;275(50):39529-42. doi: 10.1074/jbc.M005136200. J Biol Chem. 2000. PMID: 10924516
-
Glutamate dehydrogenases: the why and how of coenzyme specificity.Neurochem Res. 2014;39(3):426-32. doi: 10.1007/s11064-013-1089-x. Epub 2013 Jun 13. Neurochem Res. 2014. PMID: 23761034 Review.
-
Bovine liver glutamate dehydrogenase.Adv Protein Chem. 1976;30:101-81. doi: 10.1016/s0065-3233(08)60479-9. Adv Protein Chem. 1976. PMID: 7109 Review. No abstract available.
Cited by
-
Structural basis for leucine-induced allosteric activation of glutamate dehydrogenase.J Biol Chem. 2011 Oct 28;286(43):37406-13. doi: 10.1074/jbc.M111.260265. Epub 2011 Sep 6. J Biol Chem. 2011. PMID: 21900230 Free PMC article.
-
An enzymatic bridge between carbohydrate and amino acid metabolism: regulation of glutamate dehydrogenase by reversible phosphorylation in a severe hypoxia-tolerant crayfish.J Comp Physiol B. 2012 Apr;182(3):331-40. doi: 10.1007/s00360-011-0629-4. Epub 2011 Nov 11. J Comp Physiol B. 2012. PMID: 22076534
-
Glutamate dehydrogenase and glutamine synthetase are regulated in response to nitrogen availability in Myocbacterium smegmatis.BMC Microbiol. 2010 May 11;10:138. doi: 10.1186/1471-2180-10-138. BMC Microbiol. 2010. PMID: 20459763 Free PMC article.
-
The role of glutamine oxoglutarate aminotransferase and glutamate dehydrogenase in nitrogen metabolism in Mycobacterium bovis BCG.PLoS One. 2013 Dec 19;8(12):e84452. doi: 10.1371/journal.pone.0084452. eCollection 2013. PLoS One. 2013. PMID: 24367660 Free PMC article.
-
Unraveling nitrogen metabolism, cold and stress adaptation in polar Bosea sp. PAMC26642 through comparative genome analysis.Front Microbiol. 2025 Jan 24;15:1505699. doi: 10.3389/fmicb.2024.1505699. eCollection 2024. Front Microbiol. 2025. PMID: 39925882 Free PMC article.
References
-
- Baker, P. J., K. L. Britton, P. C. Engel, G. W. Farrants, K. S. Lilley, D. W. Rice, and T. J. Stillman. 1992. Subunit assembly and active site location in the structure of glutamate dehydrogenase. Proteins 12:75-86. - PubMed
-
- Bhuiya, M. W., H. Sakuraba, C. Kujo, N. Nunoura-Kominato, Y. Kawarabayasi, H. Kikuchi, and T. Ohshima. 2000. Glutamate dehydrogenase from the aerobic hyperthermophilic archaeon Aeropyrum pernix K1: enzymatic characterization, identification of the encoding gene, and phylogenetic implications. Extremophiles 4:333-341. - PubMed
-
- Bhuiya, M. W., H. Sakuraba, T. Ohshima, T. Imagawa, N. Katunuma, and H. Tsuge. 2005. The first crystal structure of hyperthermostable NAD-dependent glutamate dehydrogenase from Pyrobaculum islandicum. J. Mol. Biol. 345:325-337. - PubMed
-
- Bradford, M. M. 1976. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72:248-254. - PubMed
-
- Britton, K. L., P. J. Baker, D. W. Rice, and T. J. Stillman. 1992. Structural relationship between the hexameric and tetrameric family of glutamate dehydrogenases. Eur. J. Biochem. 209:851-859. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

