A general framework for designing and validating oligomer-based DNA microarrays and its application to Clostridium acetobutylicum
- PMID: 17526797
- PMCID: PMC1932840
- DOI: 10.1128/AEM.00144-07
A general framework for designing and validating oligomer-based DNA microarrays and its application to Clostridium acetobutylicum
Abstract
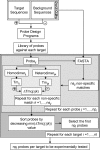
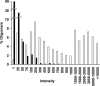
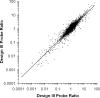
While DNA microarray analysis is widely accepted as an essential tool for modern biology, its use still eludes many researchers for several reasons, especially when microarrays are not commercially available. In that case, the design, construction, and use of microarrays for a sequenced organism constitute substantial, time-consuming, and expensive tasks. Recently, it has become possible to construct custom microarrays using industrial manufacturing processes, which offer several advantages, including speed of manufacturing, quality control, no up-front setup costs, and need-based microarray ordering. Here, we describe a strategy for designing and validating DNA microarrays manufactured using a commercial process. The 22K microarrays for the solvent producer Clostridium acetobutylicum ATCC 824 are based on in situ-synthesized 60-mers employing the Agilent technology. The strategy involves designing a large library of possible oligomer probes for each target (i.e., gene or DNA sequence) and experimentally testing and selecting the best probes for each target. The degenerate C. acetobutylicum strain M5 lacking the pSOL1 megaplasmid (with 178 annotated open reading frames [genes]) was used to estimate the level of probe cross-hybridization in the new microarrays and to establish the minimum intensity for a gene to be considered expressed. Results obtained using this microarray design were consistent with previously reported results from spotted cDNA-based microarrays. The proposed strategy is applicable to any sequenced organism.
Figures

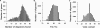

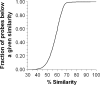
References
-
- Alsaker, K. V., C. J. Paredes, and E. T. Papoutsakis. 2005. Design, optimization and validation of genomic DNA microarrays for examining the Clostridium acetobutylicum transcriptome. Biotechnol. Bioprocess Eng. 10:432-443.
-
- Casci, T. 2001. Technology. ChIP on chips. Nat. Rev. Genet. 2:88. - PubMed
-
- Chou, H. H., A. P. Hsia, D. L. Mooney, and P. S. Schnable. 2004. PICKY: oligo microarray design for large genomes. Bioinformatics 20:2893-2902. - PubMed
-
- Churchill, G. A. 2002. Fundamentals of experimental design for cDNA microarrays. Nat. Genet. 32:490-495. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

