Comparative analysis of splice site regions by information content
- PMID: 17531798
- PMCID: PMC5054069
- DOI: 10.1016/S1672-0229(07)60003-5
Comparative analysis of splice site regions by information content
Abstract
We have applied concepts from information theory for a comparative analysis of donor (gt) and acceptor (ag) splice site regions in the genes of five different organisms by calculating their mutual information content (relative entropy) over a selected block of nucleotides. A similar pattern that the information content decreases as the block size increases was observed for both regions in all the organisms studied. This result suggests that the information required for splicing might be contained in the consensus of approximately 6-8 nt at both regions. We assume from our study that even though the nucleotides are showing some degrees of conservation in the flanking regions of the splice sites, certain level of variability is still tolerated, which leads the splicing process to occur normally even if the extent of base pairing is not fully satisfied. We also suggest that this variability can be compensated by recognizing different splice sites with different spliceosomal factors.
Figures

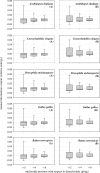
Similar articles
-
Compensatory relationship between splice sites and exonic splicing signals depending on the length of vertebrate introns.BMC Genomics. 2006 Dec 8;7:311. doi: 10.1186/1471-2164-7-311. BMC Genomics. 2006. PMID: 17156453 Free PMC article.
-
Challenging the spliceosome machine.Genome Biol. 2006;7(1):R3. doi: 10.1186/gb-2006-7-1-r3. Epub 2006 Jan 17. Genome Biol. 2006. PMID: 16507135 Free PMC article.
-
Why Selection Might Be Stronger When Populations Are Small: Intron Size and Density Predict within and between-Species Usage of Exonic Splice Associated cis-Motifs.Mol Biol Evol. 2015 Jul;32(7):1847-61. doi: 10.1093/molbev/msv069. Epub 2015 Mar 13. Mol Biol Evol. 2015. PMID: 25771198 Free PMC article.
-
Exonization of transposed elements: A challenge and opportunity for evolution.Biochimie. 2011 Nov;93(11):1928-34. doi: 10.1016/j.biochi.2011.07.014. Epub 2011 Jul 26. Biochimie. 2011. PMID: 21787833 Review.
-
Latent splice sites and stop codons revisited.RNA. 2004 Jan;10(1):5-6. doi: 10.1261/rna.5211704. RNA. 2004. PMID: 14681578 Free PMC article. Review.
References
-
- Lewin B. Genes VII. Oxford University Press; New York, USA: 2000. Nuclear splicing.
-
- Brunak S. Prediction of human mRNA donor and acceptor sites from the DNA sequence. J. Mol. Biol. 1991;220:49–65. - PubMed
-
- Chen T. Prediction of splice sites with dependency graphs and their expanded Bayesian networks. Bioinformatics. 2005;21:471–482. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

