Mastermind-1 is required for Notch signal-dependent steps in lymphocyte development in vivo
- PMID: 17535917
- PMCID: PMC1887568
- DOI: 10.1073/pnas.0700240104
Mastermind-1 is required for Notch signal-dependent steps in lymphocyte development in vivo
Abstract
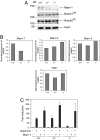
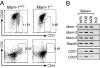
Mastermind (Mam) is one of the elements of Notch signaling, an ancient system that plays a pivotal role in metazoan development. Genetic analyses in Drosophila and Caenorhabditis elegans have shown Mam to be an essential positive regulator of this signaling pathway in these species. Mam proteins bind to and stabilize the DNA-binding complex of the intracellular domains of Notch and CBF-1, Su(H), Lag-1 (CSL) DNA-binding proteins in the nucleus. Mammals have three Mam proteins, which show remarkable similarities in their functions while having an unusual structural diversity. There have also been recent indications that Mam-1 functionally interacts with other transcription factors including p53 tumor suppressor. We herein describe that Mam-1 deficiency in mice abolishes the development of splenic marginal zone B cells, a subset strictly dependent on Notch2, a CSL protein and Delta1 ligand. Mam-1 deficiency also causes a partially impaired development of early thymocytes, while not affecting the generation of definitive hematopoiesis, processes that are dependent on Notch1. We also demonstrate the transcriptional activation of a target promoter by constitutively active forms of Notch to decrease severalfold in cultured Mam-1-deficient cells. These results indicate that Mam-1 is thus required to some extent for Notch-dependent stages in lymphopoiesis, thus supporting the notion that Mam is an essential component of the canonical Notch pathway in mammals.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
-
- Artavanis-Tsakonas S, Rand MD, Lake RJ. Science. 1999;284:770–776. - PubMed
-
- Weng AP, Ferrando AA, Lee W, Morris JP, IV, Silverman LB, Sanchez-Irizarry C, Blacklow SC, Look AT, Aster JC. Science. 2004;306:269–271. - PubMed
-
- Garg V, Muth AN, Ransom JF, Schluterman MK, Barnes R, King IN, Grossfeld PD, Srivastava D. Nature. 2005;437:270–274. - PubMed
-
- Mumm JS, Kopan R. Dev Biol. 2000;228:151–165. - PubMed
-
- Kumano K, Chiba S, Kunisato A, Sata M, Saito T, Nakagami-Yamaguchi E, Yamaguchi T, Masuda S, Shimizu K, Takahashi T, et al. Immunity. 2003;18:699–711. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

