CRISPRFinder: a web tool to identify clustered regularly interspaced short palindromic repeats
- PMID: 17537822
- PMCID: PMC1933234
- DOI: 10.1093/nar/gkm360
CRISPRFinder: a web tool to identify clustered regularly interspaced short palindromic repeats
Abstract
Clustered regularly interspaced short palindromic repeats (CRISPRs) constitute a particular family of tandem repeats found in a wide range of prokaryotic genomes (half of eubacteria and almost all archaea). They consist of a succession of highly conserved regions (DR) varying in size from 23 to 47 bp, separated by similarly sized unique sequences (spacer) of usually viral origin. A CRISPR cluster is flanked on one side by an AT-rich sequence called the leader and assumed to be a transcriptional promoter. Recent studies suggest that this structure represents a putative RNA-interference-based immune system. Here we describe CRISPRFinder, a web service offering tools to (i) detect CRISPRs including the shortest ones (one or two motifs); (ii) define DRs and extract spacers; (iii) get the flanking sequences to determine the leader; (iv) blast spacers against Genbank database and (v) check if the DR is found elsewhere in prokaryotic sequenced genomes. CRISPRFinder is freely accessible at http://crispr.u-psud.fr/Server/CRISPRfinder.php.
Figures
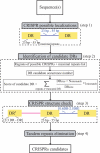

References
-
- Mojica FJ, Ferrer C, Juez G, Rodriguez-Valera F. Long stretches of short tandem repeats are present in the largest replicons of the Archaea Haloferax mediterranei and Haloferax volcanii and could be involved in replicon partitioning. Mol. Microbiol. 1995;17:85–93. - PubMed
-
- Mojica FJ, Diez-Villasenor C, Soria E, Juez G. Biological significance of a family of regularly spaced repeats in the genomes of Archaea, Bacteria and mitochondria. Mol. Microbiol. 2000;36:244–246. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

