Forced selection of tRNA(Glu) reveals the importance of two adenosine-rich RNA loops within the U5-PBS for SIV(smmPBj) replication
- PMID: 17543363
- PMCID: PMC2040306
- DOI: 10.1016/j.virol.2007.04.025
Forced selection of tRNA(Glu) reveals the importance of two adenosine-rich RNA loops within the U5-PBS for SIV(smmPBj) replication
Abstract
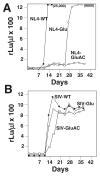
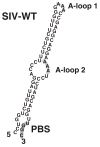
Simian immunodeficiency virus (SIV) and human immunodeficiency virus (HIV-1) preferentially select and use tRNA(Lys,3) as the primer for initiation of reverse transcription. Previous studies have shown that HIV-1 can be forced to use tRNA(Glu) if mutations are made within the primer-binding site (PBS) and a region upstream, A-loop, to be complementary to the 3'-terminal 18 nucleotides and anticodon loop of tRNA(Glu). To examine the primer preference of SIV, mutations were made within the PBS of SIV(smmPBj) to be complementary to tRNA(Glu). Analysis of the production of infectious virus revealed that SIV(smmPBj) with the PBS complementary to tRNA(Glu) retained approximately 80% infectivity of the wild type. However, modification of the U5 of SIV(smmPBj) to alter nucleotides to be complementary to the anticodon of tRNA(Glu), in combination with the PBS complementary to tRNA(Glu), drastically reduced the production of infectious SIV(smmPBj) to less than 1% that of wild type. The replication of SIV(smmPBj) with the PBS complementary to tRNA(Glu) was similar to that of the wild type virus, while the replication of SIV(smmPBj) with PBS and A-loop complementary to tRNA(Glu) was delayed compared to that of wild type virus. Analysis of the PBS regions revealed that the virus with the PBS complementary to tRNA(Glu) reverted quickly, within 4 days, to be complementary to tRNA(Lys,3), while the virus with PBS and A-loop complementary to tRNA(Glu) retained the PBS for a longer time during in vitro culture although following extended replication both the A-loop and PBS of SIV(smmPBj) reverted to be complementary to tRNA(Lys,3). RNA modeling of SIV(smmPBj) U5-PBS by m-fold revealed two potential A-loop regions. Mutations in either A-loop drastically effected replication in human PBMC. Analysis of the A-loops following in vitro replication revealed that both reverted to the wild type sequence. The results of these studies demonstrate that SIV(smmPBj), like HIV-1, preferentially utilizes tRNA(Lys,3) as a primer for reverse transcription for high level replication, but unlike HIV-1 selection may involve the use of two adenosine-rich loops.
Figures

Similar articles
-
Murine leukemia virus with a primer-binding site complementary to tRNALys,3 adapts to select new tRNAs for replication following extended in vitro culture.Virology. 2005 Mar 15;333(2):337-48. doi: 10.1016/j.virol.2004.12.033. Virology. 2005. PMID: 15721366
-
Probing the importance of tRNA anticodon: human immunodeficiency virus type 1 (HIV-1) RNA genome complementarity with an HIV-1 that selects tRNA(Glu) for replication.J Virol. 2003 Aug;77(16):8756-64. doi: 10.1128/jvi.77.16.8756-8764.2003. J Virol. 2003. PMID: 12885895 Free PMC article.
-
The G490E mutation in reverse transcriptase does not impact tRNA primer selection by HIV-1 with altered PBS and A-loop.Virology. 2006 Sep 1;352(2):380-9. doi: 10.1016/j.virol.2006.04.019. Epub 2006 Jun 15. Virology. 2006. PMID: 16781756
-
HIV-1 reverse transcription initiation: a potential target for novel antivirals?Virus Res. 2008 Jun;134(1-2):4-18. doi: 10.1016/j.virusres.2007.12.009. Epub 2008 Feb 6. Virus Res. 2008. PMID: 18255184 Review.
-
Initiation of HIV-1 reverse transcription and functional role of nucleocapsid-mediated tRNA/viral genome interactions.Virus Res. 2012 Nov;169(2):324-39. doi: 10.1016/j.virusres.2012.06.006. Epub 2012 Jun 18. Virus Res. 2012. PMID: 22721779 Review.
Cited by
-
Reverse Transcription of Retroviruses and LTR Retrotransposons.Microbiol Spectr. 2015 Apr;3(2):MDNA3-0027-2014. doi: 10.1128/microbiolspec.MDNA3-0027-2014. Microbiol Spectr. 2015. PMID: 26104704 Free PMC article. Review.
References
-
- Abraham KE, McMillen D, Brewer KL. The effects of endogenous interleukin-10 on gray matter damage and the development of pain behaviors following ecitotoxic spinal cord injury in the mouse. Neuroscience. 2004;124(4):945–952. - PubMed
-
- Baltimore D. RNA-dependent DNA polymerase in virions of RNA tumour viruses. Nature. 1970;226(5252):1209–1211. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

