Structure and function of cold shock proteins in archaea
- PMID: 17545280
- PMCID: PMC1951829
- DOI: 10.1128/JB.00395-07
Structure and function of cold shock proteins in archaea
Abstract
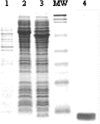
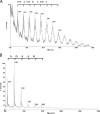
Archaea are abundant and drive critical microbial processes in the Earth's cold biosphere. Despite this, not enough is known about the molecular mechanisms of cold adaptation and no biochemical studies have been performed on stenopsychrophilic archaea (e.g., Methanogenium frigidum). This study examined the structural and functional properties of cold shock proteins (Csps) from archaea, including biochemical analysis of the Csp from M. frigidum. csp genes are present in most bacteria and some eucarya but absent from most archaeal genome sequences, most notably, those of all archaeal thermophiles and hyperthermophiles. In bacteria, Csps are small, nucleic acid binding proteins involved in a variety of cellular processes, such as transcription. In this study, archaeal Csp function was assessed by examining the ability of csp genes from psychrophilic and mesophilic Euryarchaeota and Crenarchaeota to complement a cold-sensitive growth defect in Escherichia coli. In addition, an archaeal gene with a cold shock domain (CSD) fold but little sequence identity to Csps was also examined. Genes encoding Csps or a CSD structural analog from three psychrophilic archaea rescued the E. coli growth defect. The three proteins were predicted to have a higher content of solvent-exposed basic residues than the noncomplementing proteins, and the basic residues were located on the nucleic acid binding surface, similar to their arrangement in E. coli CspA. The M. frigidum Csp was purified and found to be a single-domain protein that folds by a reversible two-state mechanism and to exhibit a low conformational stability typical of cold-adapted proteins. Moreover, M. frigidum Csp was characterized as binding E. coli single-stranded RNA, consistent with its ability to complement function in E. coli. The studies show that some Csp and CSD fold proteins have retained sufficient similarity throughout evolution in the Archaea to be able to function effectively in the Bacteria and that the function of the archaeal proteins relates to cold adaptation. The initial biochemical analysis of M. frigidum Csp has developed a platform for further characterization and demonstrates the potential for expanding molecular studies of proteins from this important archaeal stenopsychrophile.
Figures





References
-
- Bae, W., S. Phadtare, K. Severinov, and M. Inouye. 1999. Characterization of Escherichia coli cspE, whose product negatively regulates transcription of cspA, the gene for the major cold shock protein. Mol. Microbiol. 31:1429-1441. - PubMed
-
- Béjà, O., E. V. Koonin, L. Aravind, L. T. Taylor, H. Seitz, J. L. Stein, D. C. Bensen, R. A. Feldman, R. V. Swanson, and E. F. DeLong. 2002. Comparative genomic analysis of archaeal genotypic variants in a single population and in two different oceanic provinces. Appl. Environ. Microbiol. 68:335-345. - PMC - PubMed
-
- Berquist, B. R., J. Soneja, and S. DasSarma. 2005. Comparative genomic survey of information transfer systems in two diverse extremely halophilic archaea, Halobacterium sp. strain NRC-1 and Haloarcula marismortui, p. 148-182. In N. Gunde-Cimerman, A. Oren, and A. Plemenitas (ed.), Adaptation to life at high salt concentrations in Archaea, Bacteria, and Eukarya. Springer, Dordrecht, The Netherlands.
-
- Bradford, M. M. 1976. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72:248-254. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

