Genome sequencing reveals complex secondary metabolome in the marine actinomycete Salinispora tropica
- PMID: 17563368
- PMCID: PMC1965521
- DOI: 10.1073/pnas.0700962104
Genome sequencing reveals complex secondary metabolome in the marine actinomycete Salinispora tropica
Abstract
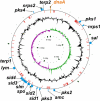
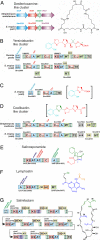
Recent fermentation studies have identified actinomycetes of the marine-dwelling genus Salinispora as prolific natural product producers. To further evaluate their biosynthetic potential, we sequenced the 5,183,331-bp S. tropica CNB-440 circular genome and analyzed all identifiable secondary natural product gene clusters. Our analysis shows that S. tropica dedicates a large percentage of its genome ( approximately 9.9%) to natural product assembly, which is greater than previous Streptomyces genome sequences as well as other natural product-producing actinomycetes. The S. tropica genome features polyketide synthase systems of every known formally classified family, nonribosomal peptide synthetases, and several hybrid clusters. Although a few clusters appear to encode molecules previously identified in Streptomyces species, the majority of the 17 biosynthetic loci are novel. Specific chemical information about putative and observed natural product molecules is presented and discussed. In addition, our bioinformatic analysis not only was critical for the structure elucidation of the polyene macrolactam salinilactam A, but its structural analysis aided the genome assembly of the highly repetitive slm loci. This study firmly establishes the genus Salinispora as a rich source of drug-like molecules and importantly reveals the powerful interplay between genomic analysis and traditional natural product isolation studies.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

