The localization of inner centromeric protein (INCENP) at the cleavage furrow is dependent on Kif12 and involves interactions of the N terminus of INCENP with the actin cytoskeleton
- PMID: 17567958
- PMCID: PMC1951774
- DOI: 10.1091/mbc.e06-10-0895
The localization of inner centromeric protein (INCENP) at the cleavage furrow is dependent on Kif12 and involves interactions of the N terminus of INCENP with the actin cytoskeleton
Abstract
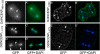
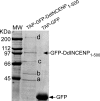
The inner centromeric protein (INCENP) and other chromosomal passenger proteins are known to localize on the cleavage furrow and to play a role in cytokinesis. However, it is not known how INCENP localizes on the furrow or whether this localization is separable from that at the midbody. Here, we show that the association of Dictyostelium INCENP (DdINCENP) with the cortex of the cleavage furrow involves interactions with the actin cytoskeleton and depends on the presence of the kinesin-6-related protein Kif12. We found that Kif12 is found on the central spindle and the cleavage furrow during cytokinesis. Kif12 is not required for the redistribution of DdINCENP from centromeres to the central spindle. However, in the absence of Kif12, DdINCENP fails to localize on the cleavage furrow. Domain analysis indicates that the N terminus of DdINCENP is necessary and sufficient for furrow localization and that it binds directly to the actin cytoskeleton. Our data suggest that INCENP moves from the central spindle to the furrow of a dividing cell by a Kif12-dependent pathway. Once INCENP reaches the equatorial cortex, it associates with the actin cytoskeleton where it then concentrates toward the end of cytokinesis.
Figures






References
-
- Adams R. R., Carmena M., Earnshaw W. C. Chromosomal passengers and the (aurora) ABCs of mitosis. Trends Cell Biol. 2001a;11:49–54. - PubMed
-
- Adams R. R., Wheatleya S. P., Gouldsworthy A. M., Kandels-Lewis S. E., Carmena M., Smythe C., Gerloff D. L., Earnshaw W. C. INCENP binds the Aurora-related kinase AIRK2 and is required to target it to chromosomes, the central spindle and cleavage furrow. Curr. Biol. 2000;10:1075–1078. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

