Identification of higher-order functional domains in the human ENCODE regions
- PMID: 17568007
- PMCID: PMC1891350
- DOI: 10.1101/gr.6081407
Identification of higher-order functional domains in the human ENCODE regions
Abstract
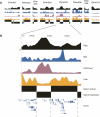
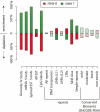
It has long been posited that human and other large genomes are organized into higher-order (i.e., greater than gene-sized) functional domains. We hypothesized that diverse experimental data types generated by The ENCODE Project Consortium could be combined to delineate active and quiescent or repressed functional domains and thereby illuminate the higher-order functional architecture of the genome. To address this, we coupled wavelet analysis with hidden Markov models for unbiased discovery of "domain-level" behavior in high-resolution functional genomic data, including activating and repressive histone modifications, RNA output, and DNA replication timing. We find that higher-order patterns in these data types are largely concordant and may be analyzed collectively in the context of HeLa cells to delineate 53 active and 62 repressed functional domains within the ENCODE regions. Active domains comprise approximately 44% of the ENCODE regions but contain approximately 75%-80% of annotated genes, transcripts, and CpG islands. Repressed domains are enriched in certain classes of repetitive elements and, surprisingly, in evolutionarily conserved nonexonic sequences. The functional domain structure of the ENCODE regions appears to be largely stable across different cell types. Taken together, our results suggest that higher-order functional domains represent a fundamental organizing principle of human genome architecture.
Figures

References
-
- Allen T.E., Herrgrd M.J., Liu M., Qiu Y., Glasner J.D., Blattner F.R., Palsson B.O., Herrgrd M.J., Liu M., Qiu Y., Glasner J.D., Blattner F.R., Palsson B.O., Liu M., Qiu Y., Glasner J.D., Blattner F.R., Palsson B.O., Qiu Y., Glasner J.D., Blattner F.R., Palsson B.O., Glasner J.D., Blattner F.R., Palsson B.O., Blattner F.R., Palsson B.O., Palsson B.O. Genome-scale analysis of the uses of the Escherichia coli genome: Model-driven analysis of heterogeneous data sets. J. Bacteriol. 2003;185:6392–6399. - PMC - PubMed
-
- Ashburner M., Ball C.A., Blake J.A., Botstein D., Butler H., Cherry J.M., Davis A.P., Dolinski K., Dwight S.S., Eppig J.T., Ball C.A., Blake J.A., Botstein D., Butler H., Cherry J.M., Davis A.P., Dolinski K., Dwight S.S., Eppig J.T., Blake J.A., Botstein D., Butler H., Cherry J.M., Davis A.P., Dolinski K., Dwight S.S., Eppig J.T., Botstein D., Butler H., Cherry J.M., Davis A.P., Dolinski K., Dwight S.S., Eppig J.T., Butler H., Cherry J.M., Davis A.P., Dolinski K., Dwight S.S., Eppig J.T., Cherry J.M., Davis A.P., Dolinski K., Dwight S.S., Eppig J.T., Davis A.P., Dolinski K., Dwight S.S., Eppig J.T., Dolinski K., Dwight S.S., Eppig J.T., Dwight S.S., Eppig J.T., Eppig J.T., et al. Gene ontology: Tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 2000;25:25–29. - PMC - PubMed
-
- Azuara V Perry P., Sauer S., Spivakov M., Jorgensen H.F., John R.M., Gouti M., Casanova M., Warnes G., Merkenschlager M., Sauer S., Spivakov M., Jorgensen H.F., John R.M., Gouti M., Casanova M., Warnes G., Merkenschlager M., Spivakov M., Jorgensen H.F., John R.M., Gouti M., Casanova M., Warnes G., Merkenschlager M., Jorgensen H.F., John R.M., Gouti M., Casanova M., Warnes G., Merkenschlager M., John R.M., Gouti M., Casanova M., Warnes G., Merkenschlager M., Gouti M., Casanova M., Warnes G., Merkenschlager M., Casanova M., Warnes G., Merkenschlager M., Warnes G., Merkenschlager M., Merkenschlager M., et al. Chromatin signatures of pluripotent cell lines. Nat. Cell Biol. 2006;8:532–538. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
