Structure and function of the HIV envelope glycoprotein as entry mediator, vaccine immunogen, and target for inhibitors
- PMID: 17586312
- PMCID: PMC7111665
- DOI: 10.1016/S1054-3589(07)55002-7
Structure and function of the HIV envelope glycoprotein as entry mediator, vaccine immunogen, and target for inhibitors
Abstract
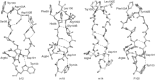
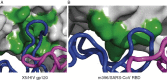
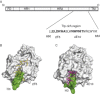
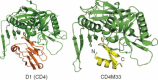
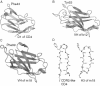
This chapter discusses the advances of the envelope glycoprotein (Env) structure as related to the interactions of conserved Env structures with receptor molecules and antibodies with implications for the design of vaccine immunogens and inhibitors. The human immunodeficiency virus (HIV) Env binds to cell surface–associated receptor (CD4) and coreceptor (CCR5 or CXCR4) by one of its two non-covalently associated subunits, gp120. The induced conformational changes activate the other subunit (gp41), which causes the fusion of the viral with the plasma cell membranes resulting in the delivery of the viral genome into the cell and the initiation of the infection cycle. As the only HIV protein exposed to the environment, the Env is also a major immunogen to which neutralizing antibodies are directed and a target that is relatively easy to access by inhibitors. A fundamental problem in the development of effective vaccines and inhibitors against HIV is the rapid generation of alterations at high levels of expression during long chronic infection and the resulting significant heterogeneity of the Env. The preservation of the Env function as an entry mediator and limitations on size and expression impose restrictions on its variability and lead to the existence of conserved structures.
Figures



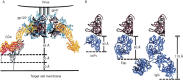
References
-
- Ahmad A., Menezes J. Antibody‐dependent cellular cytotoxicity in HIV infections. FASEB J. 1996;10:258–266. - PubMed
-
- Ahmad R., Sindhu S.T., Toma E., Morisset R., Vincelette J., Menezes J., Ahmad A. Evidence for a correlation between antibody‐dependent cellular cytotoxicity‐mediating anti‐HIV‐1 antibodies and prognostic predictors of HIV infection. J. Clin. Immunol. 2001;21:227–233. - PubMed
-
- Albert J., Abrahamsson B., Nagy K., Aurelius E., Gaines H., Nystrom G., Fenyo E.M. Rapid development of isolate‐specific neutralizing antibodies after primary HIV‐1 infection and consequent emergence of virus variants which resist neutralization by autologous sera. AIDS. 1990;4:107–112. - PubMed
-
- Alfsen A., Bomsel M. HIV‐1 gp41 envelope residues 650–685 exposed on native virus act as a lectin to bind epithelial cell galactosyl ceramide. J. Biol. Chem. 2002;277:25649–25659. - PubMed
-
- Amara R.R., Smith J.M., Staprans S.I., Montefiori D.C., Villinger F., Altman J.D., O'Neil S.P., Kozyr N.L., Xu Y., Wyatt L.S., Earl P.L., Herndon J.G. Critical role for Env as well as Gag‐Pol in control of a simian‐human immunodeficiency virus 89. 6P challenge by a DNA prime/recombinant modified vaccinia virus Ankara vaccine. J. Virol. 2002;76:6138–6146. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

