Direct localization of the 51 and 24 kDa subunits of mitochondrial complex I by three-dimensional difference imaging
- PMID: 17591445
- PMCID: PMC2700006
- DOI: 10.1016/j.jsb.2007.05.002
Direct localization of the 51 and 24 kDa subunits of mitochondrial complex I by three-dimensional difference imaging
Erratum in
- J Struct Biol. 2008 Apr;162(1):193-5
Abstract
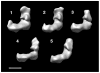
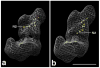
Complex I is the largest complex in the respiratory chain, and the least understood. We have determined the 3D structure of complex I from Yarrowia lipolytica lacking the flavoprotein part of the N-module, which consists of the 51 kDa (NUBM) and the 24 kDa (NUHM) subunits. The reconstruction was determined by 3D electron microscopy of single particles. A comparison to our earlier reconstruction of the complete Y. lipolytica complex I clearly assigns the two flavoprotein subunits to an outer lobe of the peripheral arm of complex I. Localizing the two subunits allowed us to fit the X-ray structure of the hydrophilic fragment of complex I from Thermus thermophilus. The fit that is most consistent with previous immuno-electron microscopic data predicts that the ubiquinone reducing catalytic center resides in the second peripheral lobe, while the 75 kDa subunit is placed near the previously seen connection between the peripheral arm and the membrane arm protrusions.
Figures





References
-
- Abdrakhmanova A, Dobrynin K, Zwicker K, Kerscher S, Brandt U. Functional sulfurtransferase is associated with mitochondrial complex I from Yarrowia lipolytica, but is not required for assembly of its iron–sulfur clusters. FEBS Letters. 2005;579:6781–6785. - PubMed
-
- Abdrakhmanova A, Zickermann V, Bostina M, Radermacher M, Schägger H, Kerscher S, Brandt U. Subunit composition of mitochondrial complex I from the yeast Yarrowia lipolytica. Biochim Biophys Acta. 2004;1658(1–2):148–56. - PubMed
-
- Belogrudov G, Hatefi Y. Catalytic sector of complex I (NADH:ubiquinone oxidoreductase): subunit stoichiometry and substrate-induced conformation changes. Biochemistry. 1994;33(15):4571–6. - PubMed
-
- Böttcher B, Scheide D, Hesterberg M, Nagel-Steger L, Friedrich T. A novel, enzymatically active conformation of the Escherichia coli ADH:ubiquinone oxidoreductase (complex I) Journal of Biological Chemistry. 2002;277(20):17970–17977. - PubMed
-
- Brandt U. Proton-translocation by membrane-bound NADH:ubiquinone-oxidoreductase (complex I) through redox-gated ligand conduction. Biochim Biophys Acta. 1997;1318(1–2):79–91. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

