Reclassification of phenotypically identified staphylococcus intermedius strains
- PMID: 17596353
- PMCID: PMC2045239
- DOI: 10.1128/JCM.00360-07
Reclassification of phenotypically identified staphylococcus intermedius strains
Abstract
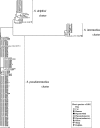
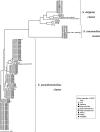
To reclassify phenotypically identified Staphylococcus intermedius strains, which might include true S. intermedius strains and novel species such as Staphylococcus pseudintermedius and Staphylococcus delphini, we analyzed molecular phylogenies and phenotypic characteristics of 117 S. intermedius group (SIG) strains tentatively identified as being S. intermedius by the Rapid ID32 Staph assay. From phylogenetic analyses of sodA and hsp60 sequences, the SIG strains were divided into three clusters, which belonged to S. pseudintermedius LMG 22219(T), S. intermedius ATCC 29663(T), and S. delphini LMG 22190(T). All the SIG strains from dogs, cats, and humans were identified as being S. pseudintermedius. The wild pigeon strains, except one, were identified as being S. intermedius, and strains from all domestic pigeons, one wild pigeon, horses, and a mink were identified as being S. delphini. In addition, a phylogenetic analysis of nuc genes revealed that S. delphini strains were divided into two clusters: one was the cluster (S. delphini group A) that belonged to S. delphini LMG 22190(T), and the other was the cluster (S. delphini group B) that was more related to S. pseudintermedius LMG 22219(T) than S. delphini LMG 22190(T). The DNA-DNA hybridization results showed that S. delphini group B strains were distinguished from S. delphini group A, S. intermedius, and S. pseudintermedius strains. S. intermedius is distinguishable from S. pseudintermedius or S. delphini by positive arginine dihydrolase and acid production from beta-gentiobiose and d-mannitol. However, phenotypical characteristics to differentiate S. delphini group A, S. delphini group B, and S. pseudintermedius were not found. In conclusion, SIG strains were reclassified into four clusters with three established and one probably novel species.
Figures

References
-
- Aarestrup, F. M. 2001. Comparative ribotyping of Staphylococcus intermedius isolated from members of the Canoidea gives possible evidence for host-specificity and co-evolution of bacteria and hosts. Int. J. Syst. Evol. Microbiol. 51:1343-1347. - PubMed
-
- Becker, K., C. von Eiff, B. Keller, M. Bruck, J. Etienne, and G. Peters. 2005. Thermonuclease gene as a target for specific identification of Staphylococcus intermedius isolates: use of a PCR-DNA enzyme immunoassay. Diagn. Microbiol. Infect. Dis. 51:237-244. - PubMed
-
- Bes, M., S. L. Saidi, F. Becharnia, H. Meugnier, F. Vandenesch, J. Etienne, and J. Freney. 2002. Population diversity of Staphylococcus intermedius isolates from various host species: typing by 16S-23S intergenic ribosomal DNA spacer polymorphism analysis. J. Clin. Microbiol. 40:2275-2277. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

