Geranylgeranyl diphosphate synthase in fission yeast is a heteromer of farnesyl diphosphate synthase (FPS), Fps1, and an FPS-like protein, Spo9, essential for sporulation
- PMID: 17596513
- PMCID: PMC1951748
- DOI: 10.1091/mbc.e07-02-0112
Geranylgeranyl diphosphate synthase in fission yeast is a heteromer of farnesyl diphosphate synthase (FPS), Fps1, and an FPS-like protein, Spo9, essential for sporulation
Abstract
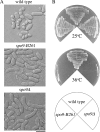
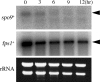
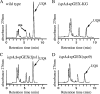
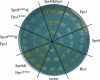
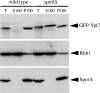
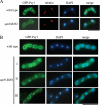
Both farnesyl diphosphate synthase (FPS) and geranylgeranyl diphosphate synthase (GGPS) are key enzymes in the synthesis of various isoprenoid-containing compounds and proteins. Here, we describe two novel Schizosaccharomyces pombe genes, fps1(+) and spo9(+), whose products are similar to FPS in primary structure, but whose functions differ from one another. Fps1 is essential for vegetative growth, whereas, a spo9 null mutant exhibits temperature-sensitive growth. Expression of fps1(+), but not spo9(+), suppresses the lethality of a Saccharomyces cerevisiae FPS-deficient mutant and also restores ubiquinone synthesis in an Escherichia coli ispA mutant, which lacks FPS activity, indicating that S. pombe Fps1 in fact functions as an FPS. In contrast to a typical FPS gene, no apparent GGPS homologues have been found in the S. pombe genome. Interestingly, although neither fps1(+) nor spo9(+) expression alone in E. coli confers clear GGPS activity, coexpression of both genes induces such activity. Moreover, the GGPS activity is significantly reduced in the spo9 mutant. In addition, the spo9 mutation perturbs the membrane association of a geranylgeranylated protein, but not that of a farnesylated protein. Yeast two-hybrid and coimmunoprecipitation analyses indicate that Fps1 and Spo9 physically interact. Thus, neither Fps1 nor Spo9 alone functions as a GGPS, but the two proteins together form a complex with GGPS activity. Because spo9 was originally identified as a sporulation-deficient mutant, we show here that expansion of the forespore membrane is severely inhibited in spo9Delta cells. Electron microscopy revealed significant accumulation membrane vesicles in spo9Delta cells. We suggest that lack of GGPS activity in a spo9 mutant results in impaired protein prenylation in certain proteins responsible for secretory function, thereby inhibiting forespore membrane formation.
Figures






Similar articles
-
Heteromer formation of a long-chain prenyl diphosphate synthase from fission yeast Dps1 and budding yeast Coq1.FEBS J. 2008 Jul;275(14):3653-68. doi: 10.1111/j.1742-4658.2008.06510.x. FEBS J. 2008. PMID: 18540885
-
The fission yeast spo14+ gene encoding a functional homologue of budding yeast Sec12 is required for the development of forespore membranes.Mol Biol Cell. 2003 Mar;14(3):1109-24. doi: 10.1091/mbc.e02-08-0504. Mol Biol Cell. 2003. PMID: 12631727 Free PMC article.
-
The Schizosaccharomyces pombe spo20(+) gene encoding a homologue of Saccharomyces cerevisiae Sec14 plays an important role in forespore membrane formation.Mol Biol Cell. 2001 Apr;12(4):901-17. doi: 10.1091/mbc.12.4.901. Mol Biol Cell. 2001. PMID: 11294895 Free PMC article.
-
Vesicle-mediated protein transport pathways to the vacuole in Schizosaccharomyces pombe.Cell Struct Funct. 2003 Oct;28(5):399-417. doi: 10.1247/csf.28.399. Cell Struct Funct. 2003. PMID: 14745133 Review.
-
Regulation of entry into gametogenesis.Philos Trans R Soc Lond B Biol Sci. 2011 Dec 27;366(1584):3521-31. doi: 10.1098/rstb.2011.0081. Philos Trans R Soc Lond B Biol Sci. 2011. PMID: 22084379 Free PMC article. Review.
Cited by
-
A genetic and pharmacological analysis of isoprenoid pathway by LC-MS/MS in fission yeast.PLoS One. 2012;7(11):e49004. doi: 10.1371/journal.pone.0049004. Epub 2012 Nov 7. PLoS One. 2012. PMID: 23145048 Free PMC article.
-
Sporulation: A response to starvation in the fission yeast Schizosaccharomyces pombe.Microbiologyopen. 2022 Jun;11(3):e1303. doi: 10.1002/mbo3.1303. Microbiologyopen. 2022. PMID: 35765188 Free PMC article. Review.
-
Structure of a heterotetrameric geranyl pyrophosphate synthase from mint (Mentha piperita) reveals intersubunit regulation.Plant Cell. 2010 Feb;22(2):454-67. doi: 10.1105/tpc.109.071738. Epub 2010 Feb 5. Plant Cell. 2010. PMID: 20139160 Free PMC article.
-
Short-chain isoprenyl diphosphate synthases of lavender (Lavandula).Plant Mol Biol. 2020 Mar;102(4-5):517-535. doi: 10.1007/s11103-020-00962-8. Epub 2020 Jan 11. Plant Mol Biol. 2020. PMID: 31927660
-
Structure and mechanism of an Arabidopsis medium/long-chain-length prenyl pyrophosphate synthase.Plant Physiol. 2011 Mar;155(3):1079-90. doi: 10.1104/pp.110.168799. Epub 2011 Jan 10. Plant Physiol. 2011. PMID: 21220764 Free PMC article.
References
-
- Blanchard L., Karst F. Characterization of a lysine-to-glutamic acid mutation in a conservative sequence of farnesyl diphosphate synthase from Saccharomyces cerevisiae. Gene. 1993;125:185–189. - PubMed
-
- Bresch C., Muller G., Egel R. Genes involved in meiosis and sporulation of a yeast. Mol. Gen. Genet. 1968;102:301–306. - PubMed
-
- Casey P. J. Biochemistry of protein prenylation. J. Lipid Res. 1992;33:1731–1740. - PubMed
-
- Casey P. J. Lipid modifications of G proteins. Curr. Opin. Cell Biol. 1994;6:219–225. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

