Paradigm development: comparative and predictive 3D modeling of HIV-1 Virion Infectivity Factor (Vif)
- PMID: 17597910
- PMCID: PMC1891711
- DOI: 10.6026/97320630001290
Paradigm development: comparative and predictive 3D modeling of HIV-1 Virion Infectivity Factor (Vif)
Abstract
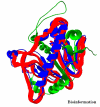
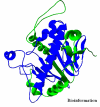
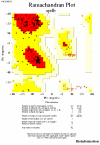
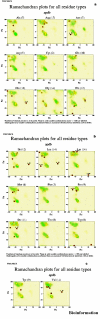
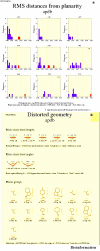
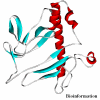
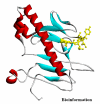
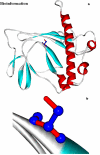
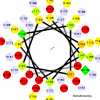
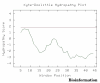
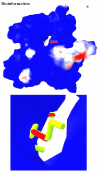
Obtaining structural information about Vif is of interest for several reasons that include the study of the interaction of Vif with APOBEC3G, a resistance factor. Vif is a potential drug target and its function is essential for the HIV-1 infectivity process. To study Vif mechanism of action, we need to decipher its structure. Pivotal in this approach is the painstaking prediction of its protein structure. The three-dimensional (3D) crystal structure for Vif has not been established. In order to understand its mechanism of action, information on the structure of Vif is very much needed. Therefore we undertook this study based on the hypothesis that information from structurally homologous proteins can be used to predict the 3D structure of Vif by computer modeling and threading. As a result the structure of HIV-1 Vif has been modeled and deposited in the theoretical models section and accepted with the PDB code 1VZF. Here, we present the results of the comparative modeling strategy we used to predict the 3D structure of Vif.
Figures


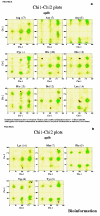
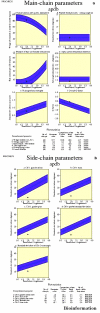
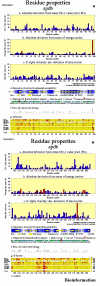
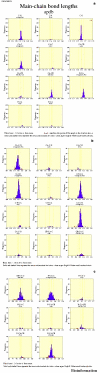




Similar articles
-
Cellular interactions of virion infectivity factor (Vif) as potential therapeutic targets: APOBEC3G and more?Curr Drug Targets. 2006 Dec;7(12):1583-93. doi: 10.2174/138945006779025356. Curr Drug Targets. 2006. PMID: 17168833 Review.
-
Influence of primate lentiviral Vif and proteasome inhibitors on human immunodeficiency virus type 1 virion packaging of APOBEC3G.J Virol. 2004 Feb;78(4):2072-81. doi: 10.1128/jvi.78.4.2072-2081.2004. J Virol. 2004. PMID: 14747572 Free PMC article.
-
Species-specific exclusion of APOBEC3G from HIV-1 virions by Vif.Cell. 2003 Jul 11;114(1):21-31. doi: 10.1016/s0092-8674(03)00515-4. Cell. 2003. PMID: 12859895
-
A single amino acid of APOBEC3G controls its species-specific interaction with virion infectivity factor (Vif).Proc Natl Acad Sci U S A. 2004 Mar 16;101(11):3927-32. doi: 10.1073/pnas.0307132101. Epub 2004 Feb 20. Proc Natl Acad Sci U S A. 2004. PMID: 14978281 Free PMC article.
-
Viral infectivity factor: a novel therapeutic strategy to block HIV-1 replication.Mini Rev Med Chem. 2013 Jun;13(7):1047-55. doi: 10.2174/1389557511313070008. Mini Rev Med Chem. 2013. PMID: 23621690 Review.
Cited by
-
Structural analysis of viral infectivity factor of HIV type 1 and its interaction with A3G, EloC and EloB.PLoS One. 2014 Feb 26;9(2):e89116. doi: 10.1371/journal.pone.0089116. eCollection 2014. PLoS One. 2014. PMID: 24586532 Free PMC article.
-
Tumultuous relationship between the human immunodeficiency virus type 1 viral infectivity factor (Vif) and the human APOBEC-3G and APOBEC-3F restriction factors.Microbiol Mol Biol Rev. 2009 Jun;73(2):211-32. doi: 10.1128/MMBR.00040-08. Microbiol Mol Biol Rev. 2009. PMID: 19487726 Free PMC article. Review.
-
Study on cocoonase, sericin, and degumming of silk cocoon: computational and experimental.J Genet Eng Biotechnol. 2021 Feb 16;19(1):32. doi: 10.1186/s43141-021-00125-2. J Genet Eng Biotechnol. 2021. PMID: 33594479 Free PMC article.
-
Mutational analysis of the HIV-1 auxiliary protein Vif identifies independent domains important for the physical and functional interaction with HIV-1 reverse transcriptase.Nucleic Acids Res. 2009 Jun;37(11):3660-9. doi: 10.1093/nar/gkp226. Epub 2009 Apr 15. Nucleic Acids Res. 2009. PMID: 19369217 Free PMC article.
-
The dimerization domain of HIV-1 viral infectivity factor Vif is required to block virion incorporation of APOBEC3G.Retrovirology. 2007 Nov 24;4:81. doi: 10.1186/1742-4690-4-81. Retrovirology. 2007. PMID: 18036235 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
