Function of periplasmic hydrogenases in the sulfate-reducing bacterium Desulfovibrio vulgaris Hildenborough
- PMID: 17601789
- PMCID: PMC1951932
- DOI: 10.1128/JB.00747-07
Function of periplasmic hydrogenases in the sulfate-reducing bacterium Desulfovibrio vulgaris Hildenborough
Abstract
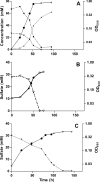
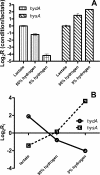
The sulfate-reducing bacterium Desulfovibrio vulgaris Hildenborough possesses four periplasmic hydrogenases to facilitate the oxidation of molecular hydrogen. These include an [Fe] hydrogenase, an [NiFeSe] hydrogenase, and two [NiFe] hydrogenases encoded by the hyd, hys, hyn1, and hyn2 genes, respectively. In order to understand their cellular functions, we have compared the growth rates of existing (hyd and hyn1) and newly constructed (hys and hyn-1 hyd) mutants to those of the wild type in defined media in which lactate or hydrogen at either 5 or 50% (vol/vol) was used as the sole electron donor for sulfate reduction. Only strains missing the [Fe] hydrogenase were significantly affected during growth with lactate or with 50% (vol/vol) hydrogen as the sole electron donor. When the cells were grown at low (5% [vol/vol]) hydrogen concentrations, those missing the [NiFeSe] hydrogenase suffered the greatest impairment. The growth rate data correlated strongly with gene expression results obtained from microarray hybridizations and real-time PCR using mRNA extracted from cells grown under the three conditions. Expression of the hys genes followed the order 5% hydrogen>50% hydrogen>lactate, whereas expression of the hyd genes followed the reverse order. These results suggest that growth with lactate and 50% hydrogen is associated with high intracellular hydrogen concentrations, which are best captured by the higher activity, lower affinity [Fe] hydrogenase. In contrast, growth with 5% hydrogen is associated with a low intracellular hydrogen concentration, requiring the lower activity, higher affinity [NiFeSe] hydrogenase.
Figures

References
-
- Alexeyev, M. F., and I. N. Shokolenko. 1995. Mini-Tn10 transposon derivatives for insertion mutagenesis and gene delivery into the chromosome of gram negative bacteria. Gene 160:59-62. - PubMed
-
- Bender, K. S., H.-C. Yen, and J. D. Wall. 2006. Analysing the metabolic capabilities of Desulfovibrio species through genetic manipulation. Biotechnol. Genet. Eng. Rev. 23:157-174. - PubMed
-
- Dolla, A., B. K. Pohorelic, J. K. Voordouw, and G. Voordouw. 2000. Deletion of the hmc operon of Desulfovibrio vulgaris subsp. vulgaris Hildenborough hampers hydrogen metabolism and low-redox-potential niche establishment. Arch. Microbiol. 174:143-151. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

