Cytophobic surface modification of microfluidic arrays for in situ parallel peptide synthesis and cell adhesion assays
- PMID: 17605465
- PMCID: PMC2546499
- DOI: 10.1021/bp070070a
Cytophobic surface modification of microfluidic arrays for in situ parallel peptide synthesis and cell adhesion assays
Abstract
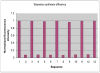
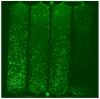
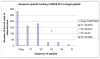
A combination of PEG-based surface passivation techniques and spatially addressable SPPS (solid-phase peptide synthesis) was used to demonstrate a highly specific cell-peptide adhesion assay on a microfluidic platform. The surface of a silicon-glass microchip was modified to form a mixed self-assembled monolayer that presented PEG moieties interspersed with reactive amino terminals. The PEG provided biomolecular inertness and the reactive amino groups were used for consequent peptide synthesis. The cytophobicity of the surface was characterized by on-chip fluorescent binding assays and was found to be resistant to nonspecific attachment of cells and proteins. An integrated system for parallel peptide synthesis on this reactive amino surface was developed using photogenerated acid chemistry and digital microlithography. A constant synthesis efficiency of >98% was observed for up to 7mer peptides. To demonstrate specific cell adhesion on these synthetic peptide arrays, variations of a 7mer cell binding peptide that binds to murine B lymphoma cells were synthesized. Sequence-specific binding was observed on incubation with fluorescently labeled, intact murine B lymphoma cells, and key residues for binding were identified by deletional analysis.
Figures
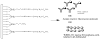
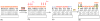

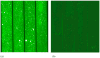


Similar articles
-
Fabrication of non-biofouling polyethylene glycol micro- and nanochannels by ultraviolet-assisted irreversible sealing.Lab Chip. 2006 Nov;6(11):1432-7. doi: 10.1039/b610503c. Epub 2006 Sep 14. Lab Chip. 2006. PMID: 17066166
-
Arrays for the combinatorial exploration of cell adhesion.J Am Chem Soc. 2004 Sep 8;126(35):10808-9. doi: 10.1021/ja0474291. J Am Chem Soc. 2004. PMID: 15339142
-
Combinatorial peptide synthesis on a microchip.Curr Protoc Protein Sci. 2009 Aug;Chapter 18:18.2.1-18.2.13. doi: 10.1002/0471140864.ps1802s57. Curr Protoc Protein Sci. 2009. PMID: 19688736
-
A comparison of gold nanoparticle surface co-functionalization approaches using Polyethylene Glycol (PEG) and the effect on stability, non-specific protein adsorption and internalization.Mater Sci Eng C Mater Biol Appl. 2016 May;62:710-8. doi: 10.1016/j.msec.2016.02.003. Epub 2016 Feb 8. Mater Sci Eng C Mater Biol Appl. 2016. PMID: 26952476
-
Tuning the Surface Interactions between Single Cells and an OSTE+ Microwell Array for Enhanced Single Cell Manipulation.ACS Appl Mater Interfaces. 2021 Jan 20;13(2):2316-2326. doi: 10.1021/acsami.0c19657. Epub 2021 Jan 7. ACS Appl Mater Interfaces. 2021. PMID: 33411502
Cited by
-
Facile construction of fluorescent peptide microarrays: One-step fluorescent derivatization of sub-microscale peptide aldehydes for selective terminal immobilization.Anal Biochem. 2010 Mar 1;398(1):132-4. doi: 10.1016/j.ab.2009.11.001. Epub 2009 Nov 10. Anal Biochem. 2010. PMID: 19903446 Free PMC article.
References
-
- Frank R. High-density synthetic peptide microarrays: Emerging tools for functional genomics and proteomics. Combinatorial Chemistry & High Throughput Screening. 2002;5(6):429–440. - PubMed
-
- Falsey JR, et al. Peptide and small molecule microarray for high throughput cell adhesion and functional assays. Bioconjugate Chemistry. 2001;12(3):346–353. - PubMed
-
- Houseman BT, et al. Peptide chips for the quantitative evaluation of protein kinase activity. Nature Biotechnology. 2002;20(3):270–274. - PubMed
-
- Panicker RC, Huang X, Yao SQ. Recent advances in peptide-based microarray technologies. Combinatorial Chemistry & High Throughput Screening. 2004;7(6):547–556. - PubMed
-
- Min DH, Mrksich M. Peptide arrays: towards routine implementation. Current Opinion in Chemical Biology. 2004;8(5):554–558. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

