Transcription termination factor Pcf11 limits the processivity of Pol II on an HIV provirus to repress gene expression
- PMID: 17606639
- PMCID: PMC1899470
- DOI: 10.1101/gad.1542707
Transcription termination factor Pcf11 limits the processivity of Pol II on an HIV provirus to repress gene expression
Abstract
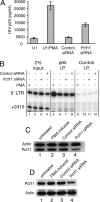
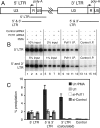
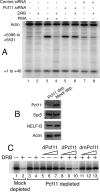
Many elongation factors in eukaryotes promote gene expression by increasing the processivity of RNA polymerase II (Pol II). However, the stability of RNA Pol II elongation complexes suggests that such complexes are not inherently prone to prematurely terminating transcription, particularly at physiological nucleotide concentrations. We show that the termination factor, Pcf11, causes premature termination on an HIV provirus. The transcription that occurs when Pcf11 is depleted from cells or an extract is no longer sensitive to 6-dichloro-1-beta-D-ribofuranosylbenzimidazole (DRB), a compound that causes premature termination. Hence, Pcf11 can act as a negative elongation factor to repress RNA Pol II gene expression in eukaryotic cells.
Figures

Similar articles
-
Human Pcf11 enhances degradation of RNA polymerase II-associated nascent RNA and transcriptional termination.Nucleic Acids Res. 2008 Feb;36(3):905-14. doi: 10.1093/nar/gkm1112. Epub 2007 Dec 17. Nucleic Acids Res. 2008. PMID: 18086705 Free PMC article.
-
Premature transcription termination complex proteins PCF11 and WDR82 silence HIV-1 expression in latently infected cells.Proc Natl Acad Sci U S A. 2023 Dec 5;120(49):e2313356120. doi: 10.1073/pnas.2313356120. Epub 2023 Nov 28. Proc Natl Acad Sci U S A. 2023. PMID: 38015843 Free PMC article.
-
Negative elongation factor (NELF) coordinates RNA polymerase II pausing, premature termination, and chromatin remodeling to regulate HIV transcription.J Biol Chem. 2013 Sep 6;288(36):25995-26003. doi: 10.1074/jbc.M113.496489. Epub 2013 Jul 24. J Biol Chem. 2013. PMID: 23884411 Free PMC article.
-
Interplay between positive and negative elongation factors: drawing a new view of DRB.Genes Cells. 1998 Jan;3(1):9-15. doi: 10.1046/j.1365-2443.1998.00162.x. Genes Cells. 1998. PMID: 9581978 Review.
-
Termination of Transcription by RNA Polymerase II: BOOM!Trends Genet. 2020 Sep;36(9):664-675. doi: 10.1016/j.tig.2020.05.008. Epub 2020 Jun 8. Trends Genet. 2020. PMID: 32527618 Review.
Cited by
-
Genome-wide mapping of yeast RNA polymerase II termination.PLoS Genet. 2014 Oct 9;10(10):e1004632. doi: 10.1371/journal.pgen.1004632. eCollection 2014 Oct. PLoS Genet. 2014. PMID: 25299594 Free PMC article.
-
Ratio-based analysis of differential mRNA processing and expression of a polyadenylation factor mutant pcfs4 using arabidopsis tiling microarray.PLoS One. 2011 Feb 25;6(2):e14719. doi: 10.1371/journal.pone.0014719. PLoS One. 2011. PMID: 21364912 Free PMC article.
-
Impact of Chromatin on HIV Replication.Genes (Basel). 2015 Sep 30;6(4):957-76. doi: 10.3390/genes6040957. Genes (Basel). 2015. PMID: 26437430 Free PMC article. Review.
-
Targeting Viral Transcription for HIV Cure Strategies.Microorganisms. 2024 Apr 8;12(4):752. doi: 10.3390/microorganisms12040752. Microorganisms. 2024. PMID: 38674696 Free PMC article. Review.
-
Promoter proximal pausing on genes in metazoans.Chromosoma. 2009 Feb;118(1):1-10. doi: 10.1007/s00412-008-0182-4. Epub 2008 Oct 2. Chromosoma. 2009. PMID: 18830703 Review.
References
-
- Aiyar S.E., Sun J.L., Blair A.L., Moskaluk C.A., Lu Y.Z., Ye Q.N., Yamaguchi Y., Mukherjee A., Ren D.M., Handa H., Sun J.L., Blair A.L., Moskaluk C.A., Lu Y.Z., Ye Q.N., Yamaguchi Y., Mukherjee A., Ren D.M., Handa H., Blair A.L., Moskaluk C.A., Lu Y.Z., Ye Q.N., Yamaguchi Y., Mukherjee A., Ren D.M., Handa H., Moskaluk C.A., Lu Y.Z., Ye Q.N., Yamaguchi Y., Mukherjee A., Ren D.M., Handa H., Lu Y.Z., Ye Q.N., Yamaguchi Y., Mukherjee A., Ren D.M., Handa H., Ye Q.N., Yamaguchi Y., Mukherjee A., Ren D.M., Handa H., Yamaguchi Y., Mukherjee A., Ren D.M., Handa H., Mukherjee A., Ren D.M., Handa H., Ren D.M., Handa H., Handa H., et al. Attenuation of estrogen receptor α-mediated transcription through estrogen-stimulated recruitment of a negative elongation factor. Genes & Dev. 2004;18:2134–2146. - PMC - PubMed
-
- Buratowski S. Connections between mRNA 3′ end processing and transcription termination. Curr. Opin. Cell Biol. 2005;17:257–261. - PubMed
-
- Cook J.A., Albacker L., August A., Henderson A.J., Albacker L., August A., Henderson A.J., August A., Henderson A.J., Henderson A.J. CD28-dependent HIV-1 transcription is associated with Vav, Rac, and NF-κ B activation. J. Biol. Chem. 2003;278:35812–35818. - PubMed
-
- Cullen B.R. The HIV-1 Tat protein: An RNA sequence-specific processivity factor. Cell. 1990;63:655–657. - PubMed
-
- Cullen B.R. Regulation of HIV-1 gene expression. FASEB J. 1991;5:2361–2368. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
