Dietary phytochemicals regulate whole-body CYP1A1 expression through an arylhydrocarbon receptor nuclear translocator-dependent system in gut
- PMID: 17607366
- PMCID: PMC1890999
- DOI: 10.1172/JCI31647
Dietary phytochemicals regulate whole-body CYP1A1 expression through an arylhydrocarbon receptor nuclear translocator-dependent system in gut
Abstract
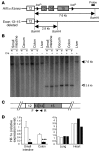
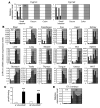
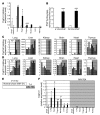
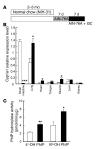
Cytochrome P450 1A1 (CYP1A1) is one of the most important detoxification enzymes due to its broad substrate specificity and wide distribution throughout the body. On the other hand, CYP1A1 can also produce highly carcinogenic intermediate metabolites through oxidation of polycyclic aromatic hydrocarbons. We describe what we believe to be a novel regulatory system for whole-body CYP1A1 expression by a factor originating in the gut. A mutant mouse was generated in which the arylhydrocarbon receptor nuclear translocator (Arnt) gene is disrupted predominantly in the gut epithelium. Surprisingly, CYP1A1 mRNA expression and enzymatic activities were markedly elevated in almost all non-gut tissues in this mouse line. The induction was even observed in early-stage embryos in pregnant mutant females. Interestingly, the upregulation was CYP1A1 selective and lost upon administration of a synthetic purified diet. Moreover, the increase was recovered by addition of the natural phytochemical indole-3-carbinol to the purified diet. These results suggest that an Arnt-dependent pathway in gut has an important role in regulation of the metabolism of dietary CYP1A1 inducers and whole-body CYP1A1 expression. This machinery might be involved in naturally occurring carcinogenic processes and/or other numerous biological responses mediated by CYP1A1 activity.
Figures

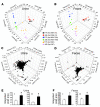
References
-
- Brooks R.A., Gooderham N.J., Edwards R.J., Boobis A.R., Winton D.J. The mutagenicity of benzo[a]pyrene in mouse small intestine. Carcinogenesis. 1999;20:109–114. - PubMed
-
- Kaminsky L.S., Zhang Q.Y. The small intestine as a xenobiotic-metabolizing organ. Drug Metab. Dispos. 2003;31:1520–1525. - PubMed
-
- Lindell M., Lang M., Lennernas H. Expression of genes encoding for drug metabolising cytochrome P450 enzymes and P-glycoprotein in the rat small intestine; comparison to the liver. Eur. J. Drug Metab. Pharmacokinet. 2003;28:41–48. - PubMed
-
- Zhang Q.Y., Dunbar D., Kaminsky L.S. Characterization of mouse small intestinal cytochrome P450 expression. Drug Metab. Dispos. 2003;31:1346–1351. - PubMed
-
- Ma Q. Induction of CYP1A1. The AhR/DRE paradigm: transcription, receptor regulation, and expanding biological roles. Curr. Drug Metab. 2001;2:149–164. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

