Structural and functional analysis of two glutamate racemase isozymes from Bacillus anthracis and implications for inhibitor design
- PMID: 17610893
- PMCID: PMC2736553
- DOI: 10.1016/j.jmb.2007.05.093
Structural and functional analysis of two glutamate racemase isozymes from Bacillus anthracis and implications for inhibitor design
Abstract
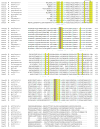
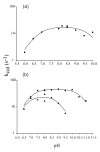
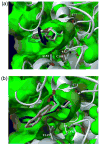
Glutamate racemase (RacE) is responsible for converting l-glutamate to d-glutamate, which is an essential component of peptidoglycan biosynthesis, and the primary constituent of the poly-gamma-d-glutamate capsule of the pathogen Bacillus anthracis. RacE enzymes are essential for bacterial growth and lack a human homolog, making them attractive targets for the design and development of antibacterial therapeutics. We have cloned, expressed and purified the two glutamate racemase isozymes, RacE1 and RacE2, from the B. anthracis genome. Through a series of steady-state kinetic studies, and based upon the ability of both RacE1 and RacE2 to catalyze the rapid formation of d-glutamate, we have determined that RacE1 and RacE2 are bona fide isozymes. The X-ray structures of B. anthracis RacE1 and RacE2, in complex with d-glutamate, were determined to resolutions of 1.75 A and 2.0 A. Both enzymes are dimers with monomers arranged in a "tail-to-tail" orientation, similar to the B. subtilis RacE structure, but differing substantially from the Aquifex pyrophilus RacE structure. The differences in quaternary structures produce differences in the active sites of racemases among the various species, which has important implications for structure-based, inhibitor design efforts within this class of enzymes. We found a Val to Ala variance at the entrance of the active site between RacE1 and RacE2, which results in the active site entrance being less sterically hindered for RacE1. Using a series of inhibitors, we show that this variance results in differences in the inhibitory activity against the two isozymes and suggest a strategy for structure-based inhibitor design to obtain broad-spectrum inhibitors for glutamate racemases.
Figures
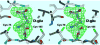


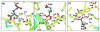


Similar articles
-
Functional comparison of the two Bacillus anthracis glutamate racemases.J Bacteriol. 2007 Jul;189(14):5265-75. doi: 10.1128/JB.00352-07. Epub 2007 May 11. J Bacteriol. 2007. PMID: 17496086 Free PMC article.
-
Glutamate Racemase Mutants of Bacillus anthracis.J Bacteriol. 2015 Jun;197(11):1854-61. doi: 10.1128/JB.00070-15. Epub 2015 Mar 16. J Bacteriol. 2015. PMID: 25777674 Free PMC article.
-
Efficient gene inactivation in Bacillus anthracis.FEMS Microbiol Lett. 2005 Apr 15;245(2):315-9. doi: 10.1016/j.femsle.2005.03.029. FEMS Microbiol Lett. 2005. PMID: 15837388
-
Glutamate racemase as a target for drug discovery.Microb Biotechnol. 2008 Sep;1(5):345-60. doi: 10.1111/j.1751-7915.2008.00031.x. Epub 2008 May 11. Microb Biotechnol. 2008. PMID: 21261855 Free PMC article. Review.
-
Biochemistry and molecular genetics of poly-gamma-glutamate synthesis.Appl Microbiol Biotechnol. 2002 Jun;59(1):9-14. doi: 10.1007/s00253-002-0984-x. Epub 2002 Apr 16. Appl Microbiol Biotechnol. 2002. PMID: 12073126 Review.
Cited by
-
Mycobacterial Targets for Thiourea Derivatives: Opportunities for Virtual Screening in Tuberculosis Drug Discovery.Curr Med Chem. 2024;31(29):4703-4724. doi: 10.2174/0109298673276076231124104513. Curr Med Chem. 2024. PMID: 38375848 Review.
-
Genome Analysis Coupled With Transcriptomics Reveals the Reduced Fitness of a Hot Spring Cyanobacterium Mastigocladus laminosus UU774 Under Exogenous Nitrogen Supplement.Front Microbiol. 2022 Jul 1;13:909289. doi: 10.3389/fmicb.2022.909289. eCollection 2022. Front Microbiol. 2022. PMID: 35847102 Free PMC article.
-
Role of purine biosynthesis in Bacillus anthracis pathogenesis and virulence.Infect Immun. 2011 Jan;79(1):153-66. doi: 10.1128/IAI.00925-10. Epub 2010 Nov 1. Infect Immun. 2011. PMID: 21041498 Free PMC article.
-
Analyses of cobalt-ligand and potassium-ligand bond lengths in metalloproteins: trends and patterns.J Mol Model. 2014 Jun;20(6):2271. doi: 10.1007/s00894-014-2271-z. Epub 2014 May 22. J Mol Model. 2014. PMID: 24850495
-
Structural basis for catalysis of a tetrameric class IIa fructose 1,6-bisphosphate aldolase from Mycobacterium tuberculosis.J Mol Biol. 2009 Mar 6;386(4):1038-53. doi: 10.1016/j.jmb.2009.01.003. Epub 2009 Jan 10. J Mol Biol. 2009. PMID: 19167403 Free PMC article.
References
-
- Atlas RM. Bioterriorism: from threat to reality. Annu Rev Microbiol. 2002;56:167–85. - PubMed
-
- Athamna A, Athamna M, Abu-Rashed N, Medlej B, Bast DJ, Rubinstein E. Selection of Bacillus anthracis isolates resistant to antibiotics. J Antimicrob Chemother. 2004;54:424–8. - PubMed
-
- Bast DJ, Athamna A, Duncan CL, de Azavedo JC, Low DE, Rahav G, Farrell D, Rubinstein E. Type II topoisomerase mutations in Bacillus anthracis associated with high-level fluoroquinolone resistance. J Antimicrob Chemother. 2004;54:90–4. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

