Characterization of the dsDNA prophage sequences in the genome of Neisseria gonorrhoeae and visualization of productive bacteriophage
- PMID: 17615066
- PMCID: PMC1931599
- DOI: 10.1186/1471-2180-7-66
Characterization of the dsDNA prophage sequences in the genome of Neisseria gonorrhoeae and visualization of productive bacteriophage
Abstract
Background: Bioinformatic analysis of the genome sequence of Neisseria gonorrhoeae revealed the presence of nine probable prophage islands. The distribution, conservation and function of many of these sequences, and their ability to produce bacteriophage particles are unknown.
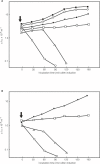
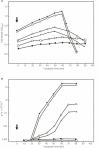
Results: Our analysis of the genomic sequence of FA1090 identified five genomic regions (NgoPhi1 - 5) that are related to dsDNA lysogenic phage. The genetic content of the dsDNA prophage sequences were examined in detail and found to contain blocks of genes encoding for proteins homologous to proteins responsible for phage DNA replication, structural proteins and proteins responsible for phage assembly. The DNA sequences from NgoPhi1, NgoPhi2 and NgoPhi3 contain some significant regions of identity. A unique region of NgoPhi2 showed very high similarity with the Pseudomonas aeruginosa generalized transducing phage F116. Comparative analysis at the nucleotide and protein levels suggests that the sequences of NgoPhi1 and NgoPhi2 encode functionally active phages, while NgoPhi3, NgoPhi4 and NgoPhi5 encode incomplete genomes. Expression of the NgoPhi1 and NgoPhi2 repressors in Escherichia coli inhibit the growth of E. coli and the propagation of phage lambda. The NgoPhi2 repressor was able to inhibit transcription of N. gonorrhoeae genes and Haemophilus influenzae HP1 phage promoters. The holin gene of NgoPhi1 (identical to that encoded by NgoPhi2), when expressed in E. coli, could serve as substitute for the phage lambda s gene. We were able to detect the presence of the DNA derived from NgoPhi1 in the cultures of N. gonorrhoeae. Electron microscopy analysis of culture supernatants revealed the presence of multiple forms of bacteriophage particles.
Conclusion: These data suggest that the genes similar to dsDNA lysogenic phage present in the gonococcus are generally conserved in this pathogen and that they are able to regulate the expression of other neisserial genes. Since phage particles were only present in culture supernatants after induction with mitomycin C, it indicates that the gonococcus also regulates the expression of bacteriophage genes.
Figures







Similar articles
-
CRISPRi-mediated repression of three cI repressors induces the expression of three related Neisseria gonorrhoeae bacteriophages.J Bacteriol. 2025 Jun 24;207(6):e0004925. doi: 10.1128/jb.00049-25. Epub 2025 May 12. J Bacteriol. 2025. PMID: 40353677 Free PMC article.
-
Analysis of the filamentous bacteriophage genomes integrated into Neisseria gonorrhoeae FA1090 chromosome.Pol J Microbiol. 2006;55(4):251-60. Pol J Microbiol. 2006. PMID: 17416061
-
Analysis of the lambdoid prophage element e14 in the E. coli K-12 genome.BMC Microbiol. 2004 Jan 20;4:4. doi: 10.1186/1471-2180-4-4. BMC Microbiol. 2004. PMID: 14733619 Free PMC article.
-
The potential for bacteriophages and prophage elements in fighting and preventing the gonorrhea.Crit Rev Microbiol. 2024 Sep;50(5):769-784. doi: 10.1080/1040841X.2023.2274849. Epub 2023 Oct 28. Crit Rev Microbiol. 2024. PMID: 37897236 Review.
-
Yet another way that phage λ manipulates its Escherichia coli host: λrexB is involved in the lysogenic-lytic switch.Mol Microbiol. 2015 May;96(4):689-93. doi: 10.1111/mmi.12969. Epub 2015 Mar 16. Mol Microbiol. 2015. PMID: 25684601 Review.
Cited by
-
CRISPRi-mediated repression of three cI repressors induces the expression of three related Neisseria gonorrhoeae bacteriophages.J Bacteriol. 2025 Jun 24;207(6):e0004925. doi: 10.1128/jb.00049-25. Epub 2025 May 12. J Bacteriol. 2025. PMID: 40353677 Free PMC article.
-
Phase-variable Type I methyltransferase M.NgoAV from Neisseria gonorrhoeae FA1090 regulates phasevarion expression and gonococcal phenotype.Front Microbiol. 2022 Oct 4;13:917639. doi: 10.3389/fmicb.2022.917639. eCollection 2022. Front Microbiol. 2022. PMID: 36267167 Free PMC article.
-
The Gonococcal Transcriptome during Infection of the Lower Genital Tract in Women.PLoS One. 2015 Aug 5;10(8):e0133982. doi: 10.1371/journal.pone.0133982. eCollection 2015. PLoS One. 2015. PMID: 26244506 Free PMC article.
-
The impact of antimicrobials on gonococcal evolution.Nat Microbiol. 2019 Nov;4(11):1941-1950. doi: 10.1038/s41564-019-0501-y. Epub 2019 Jul 29. Nat Microbiol. 2019. PMID: 31358980 Free PMC article.
-
A genetic approach to the development of new therapeutic phages to fight pseudomonas aeruginosa in wound infections.Viruses. 2012 Dec 21;5(1):15-53. doi: 10.3390/v5010015. Viruses. 2012. PMID: 23344559 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

