Yeast DNA polymerase epsilon participates in leading-strand DNA replication
- PMID: 17615360
- PMCID: PMC2233713
- DOI: 10.1126/science.1144067
Yeast DNA polymerase epsilon participates in leading-strand DNA replication
Abstract
Multiple DNA polymerases participate in replicating the leading and lagging strands of the eukaryotic nuclear genome. Although 50 years have passed since the first DNA polymerase was discovered, the identity of the major polymerase used for leading-strand replication is uncertain. We constructed a derivative of yeast DNA polymerase epsilon that retains high replication activity but has strongly reduced replication fidelity, particularly for thymine-deoxythymidine 5'-monophosphate (T-dTMP) but not adenine-deoxyadenosine 5'-monophosphate (A-dAMP) mismatches. Yeast strains with this DNA polymerase epsilon allele have elevated rates of T to A substitution mutations. The position and rate of these substitutions depend on the orientation of the mutational reporter and its location relative to origins of DNA replication and reveal a pattern indicating that DNA polymerase epsilon participates in leading-strand DNA replication.
Figures
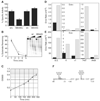

References
-
- Garg P, Burgers PM. Crit. Rev. Biochem. Mol. Biol. 2005;40:115. - PubMed
-
- Johnson A, O'Donnell M. Annu. Rev. Biochem. 2005;74:283. - PubMed
-
-
Materials and methods are available as supporting material on Science Online.
-
-
- Pavlov YI, Mian IM, Kunkel TA. Curr. Biol. 2003;13:744. - PubMed
-
- Pavlov YI, Newlon CS, Kunkel TA. Mol. Cell. 2002;10:207. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

