PAB1 self-association precludes its binding to poly(A), thereby accelerating CCR4 deadenylation in vivo
- PMID: 17620415
- PMCID: PMC1952152
- DOI: 10.1128/MCB.00734-07
PAB1 self-association precludes its binding to poly(A), thereby accelerating CCR4 deadenylation in vivo
Abstract
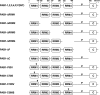
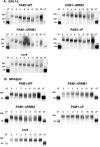
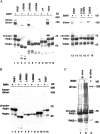
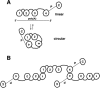
The mRNA deadenylation process, catalyzed by the CCR4 deadenylase, is known to be the major factor controlling mRNA decay rates in Saccharomyces cerevisiae. We have identified the proline-rich region and RRM1 domains of poly(A) binding protein (PAB1) as necessary for CCR4 deadenylation. Deletion of either of these regions but not other regions of PAB1 significantly reduced PAB1-PAB1 protein interactions, suggesting that PAB1 oligomerization is a required step for deadenylation. Moreover, defects in these two regions inhibited the formation of a novel, circular monomeric PAB1 species that forms in the absence of poly(A). Removal of the PAB1 RRM3 domain, which promoted PAB1 oligomerization and circularization, correspondingly accelerated CCR4 deadenylation. Circular PAB1 was unable to bind poly(A), and PAB1 multimers were severely deficient or unable to bind poly(A), implicating the PAB1 RNA binding surface as critical in making contacts that allow PAB1 self-association. These results support the model that the control of CCR4 deadenylation in vivo occurs in part through the removal of PAB1 from the poly(A) tail following its self-association into multimers and/or a circular species. Known alterations in the P domains of different PAB proteins and factors and conditions that affect PAB1 self-association would, therefore, be expected to be critical to controlling mRNA turnover in the cell.
Figures


Similar articles
-
PUF3 acceleration of deadenylation in vivo can operate independently of CCR4 activity, possibly involving effects on the PAB1-mRNP structure.J Mol Biol. 2010 Jun 18;399(4):562-75. doi: 10.1016/j.jmb.2010.04.034. Epub 2010 May 8. J Mol Biol. 2010. PMID: 20435044 Free PMC article.
-
A specific role for the C-terminal region of the Poly(A)-binding protein in mRNA decay.Nucleic Acids Res. 2007;35(18):6017-28. doi: 10.1093/nar/gkm452. Epub 2007 Aug 30. Nucleic Acids Res. 2007. PMID: 17766253 Free PMC article.
-
The RRM1 domain of the poly(A)-binding protein from Saccharomyces cerevisiae is critical to control of mRNA deadenylation.Mol Genet Genomics. 2013 Sep;288(9):401-12. doi: 10.1007/s00438-013-0759-3. Epub 2013 Jun 21. Mol Genet Genomics. 2013. PMID: 23793387 Free PMC article.
-
The Saccharomyces cerevisiae poly (A) binding protein (Pab1): Master regulator of mRNA metabolism and cell physiology.Yeast. 2019 Jan;36(1):23-34. doi: 10.1002/yea.3347. Epub 2018 Oct 17. Yeast. 2019. PMID: 30006991 Review.
-
Molecular Insights into mRNA Polyadenylation and Deadenylation.Int J Mol Sci. 2022 Sep 20;23(19):10985. doi: 10.3390/ijms231910985. Int J Mol Sci. 2022. PMID: 36232288 Free PMC article. Review.
Cited by
-
PUF3 acceleration of deadenylation in vivo can operate independently of CCR4 activity, possibly involving effects on the PAB1-mRNP structure.J Mol Biol. 2010 Jun 18;399(4):562-75. doi: 10.1016/j.jmb.2010.04.034. Epub 2010 May 8. J Mol Biol. 2010. PMID: 20435044 Free PMC article.
-
Transient CPEB dimerization and translational control.RNA. 2012 May;18(5):1050-61. doi: 10.1261/rna.031682.111. Epub 2012 Mar 28. RNA. 2012. PMID: 22456264 Free PMC article.
-
Identification of a 57S translation complex containing closed-loop factors and the 60S ribosome subunit.Sci Rep. 2018 Jul 31;8(1):11468. doi: 10.1038/s41598-018-29832-6. Sci Rep. 2018. PMID: 30065356 Free PMC article.
-
The Saccharomyces cerevisiae poly(A) binding protein Pab1 as a target for eliciting stress tolerant phenotypes.Sci Rep. 2015 Dec 14;5:18318. doi: 10.1038/srep18318. Sci Rep. 2015. PMID: 26658950 Free PMC article.
-
Interaction between the poly(A)-binding protein Pab1 and the eukaryotic release factor eRF3 regulates translation termination but not mRNA decay in Saccharomyces cerevisiae.RNA. 2015 Jan;21(1):124-34. doi: 10.1261/rna.047282.114. Epub 2014 Nov 19. RNA. 2015. PMID: 25411355 Free PMC article.
References
-
- Blanco, P., C. A. Sargent, C. A. Boucher, G. Howell, M. Ross, and N. A. Affara. 2001. A novel poly(A)-binding protein gene (PABPC5) maps to an X-specific subinterval in the Xq21.3/Yp11.2 homology block of the human sex chromosomes. Genomics 74:1-11. - PubMed
-
- Caponigro, G., and R. Parker. 1995. Multiple functions for the poly(A)-binding protein in mRNA decapping and deadenylation in yeast. Genes Dev. 9:2421-2432. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
