Recombination rate estimation in the presence of hotspots
- PMID: 17623807
- PMCID: PMC1933511
- DOI: 10.1101/gr.6386707
Recombination rate estimation in the presence of hotspots
Abstract
Fine-scale estimation of recombination rates remains a challenging problem. Experimental techniques can provide accurate estimates at fine scales but are technically challenging and cannot be applied on a genome-wide scale. An alternative source of information comes from patterns of genetic variation. Several statistical methods have been developed to estimate recombination rates from randomly sampled chromosomes. However, most such methods either make poor assumptions about recombination rate variation, or simply assume that there is no rate variation. Since the discovery of recombination hotspots, it is clear that recombination rates can vary over many orders of magnitude at the fine scale. We present a method for the estimation of recombination rates in the presence of recombination hotspots. We demonstrate that the method is able to detect and accurately quantify recombination rate heterogeneity, and is a substantial improvement over a commonly used method. We then use the method to reanalyze genetic variation data from the HLA and MS32 regions of the human genome and demonstrate that the method is able to provide accurate rate estimates and simultaneously detect hotspots.
Figures

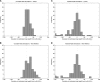
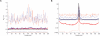
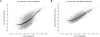
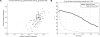

References
-
- Fearnhead P. Consistency of estimators of the population-scaled recombination rate. Theor. Popul. Biol. 2003;64:67–79. - PubMed
-
- Fearnhead P. SequenceLDhot: Detecting recombination hotspots. Bioinformatics. 2006;22:3061–3066. - PubMed
-
- Greenawalt D.M., Cui X., Wu Y., Lin Y., Wang H.Y., Luo M., Tereshchenko I.V., Hu G., Li J.Y., Chu Y., Cui X., Wu Y., Lin Y., Wang H.Y., Luo M., Tereshchenko I.V., Hu G., Li J.Y., Chu Y., Wu Y., Lin Y., Wang H.Y., Luo M., Tereshchenko I.V., Hu G., Li J.Y., Chu Y., Lin Y., Wang H.Y., Luo M., Tereshchenko I.V., Hu G., Li J.Y., Chu Y., Wang H.Y., Luo M., Tereshchenko I.V., Hu G., Li J.Y., Chu Y., Luo M., Tereshchenko I.V., Hu G., Li J.Y., Chu Y., Tereshchenko I.V., Hu G., Li J.Y., Chu Y., Hu G., Li J.Y., Chu Y., Li J.Y., Chu Y., Chu Y., et al. Strong correlation between meiotic crossovers and haplotype structure in a 2.5-Mb region on the long arm of chromosome 21. Genome Res. 2006;16:208–214. - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Research Materials
