Gfra1 silencing in mouse spermatogonial stem cells results in their differentiation via the inactivation of RET tyrosine kinase
- PMID: 17625109
- PMCID: PMC2911237
- DOI: 10.1095/biolreprod.107.062513
Gfra1 silencing in mouse spermatogonial stem cells results in their differentiation via the inactivation of RET tyrosine kinase
Abstract
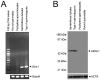
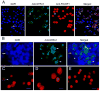
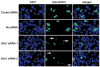
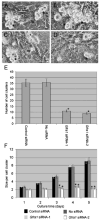
Spermatogenesis is the process by which spermatogonial stem cells divide and differentiate into sperm. The role of growth factor receptors in regulating self-renewal and differentiation of spermatogonial stem cells remains largely unclear. This study was designed to examine Gfra1 receptor expression in immature and adult mouse testes and determine the effects of Gfra1 knockdown on the proliferation and differentiation of type A spermatogonia. We demonstrated that GFRA1 was expressed in a subpopulation of spermatogonia in immature and adult mice. Neither Gfra1 mRNA nor GFRA1 protein was detected in pachytene spermatocytes and round spermatids. GFRA1 and POU5F1 (also known as OCT4), a marker for spermatogonial stem cells, were co-expressed in a subpopulation of type A spermatogonia from 6-day-old mice. In addition, the spermatogonia expressing GFRA1 exhibited a potential for proliferation and the ability to form colonies in culture, which is a characteristic of stem cells. RNA interference assays showed that Gfra1 small interfering RNAs (siRNAs) knocked down the expression of Gfra1 mRNA and GFRA1 protein in type A spermatogonia. Notably, the reduction of Gfra1 expression by Gfra1 siRNAs induced a phenotypic differentiation, as evidenced by the elevated expression of KIT, as well as the decreased expression of POU5F1 and proliferating cell nuclear antigen (PCNA). Furthermore, Gfra1 silencing resulted in a decrease in RET phosphorylation. Taken together, these data indicate that Gfra1 is expressed dominantly in mouse spermatogonial stem cells and that Gfra1 knockdown leads to their differentiation via the inactivation of RET tyrosine kinase, suggesting an essential role for Gfra1 in spermatogonial stem cell regulation.
Figures



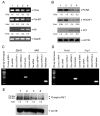
Similar articles
-
Loss of Etv5 decreases proliferation and RET levels in neonatal mouse testicular germ cells and causes an abnormal first wave of spermatogenesis.Biol Reprod. 2009 Aug;81(2):258-66. doi: 10.1095/biolreprod.108.075200. Epub 2009 Apr 15. Biol Reprod. 2009. PMID: 19369650 Free PMC article.
-
GDNF family receptor alpha1 phenotype of spermatogonial stem cells in immature mouse testes.Biol Reprod. 2005 Nov;73(5):1011-6. doi: 10.1095/biolreprod.105.043810. Epub 2005 Jul 13. Biol Reprod. 2005. PMID: 16014811
-
Glial cell-line derived neurotrophic factor-mediated RET signaling regulates spermatogonial stem cell fate.Biol Reprod. 2006 Feb;74(2):314-21. doi: 10.1095/biolreprod.105.047365. Epub 2005 Oct 19. Biol Reprod. 2006. PMID: 16237148
-
Regulation of spermatogonial stem cell self-renewal and spermatocyte meiosis by Sertoli cell signaling.Reproduction. 2015 Apr;149(4):R159-67. doi: 10.1530/REP-14-0481. Epub 2014 Dec 12. Reproduction. 2015. PMID: 25504872 Review.
-
Regulation of GDNF expression in Sertoli cells.Reproduction. 2019 Mar;157(3):R95-R107. doi: 10.1530/REP-18-0239. Reproduction. 2019. PMID: 30620720 Free PMC article. Review.
Cited by
-
Generation and characteristics of human Sertoli cell line immortalized by overexpression of human telomerase.Oncotarget. 2017 Mar 7;8(10):16553-16570. doi: 10.18632/oncotarget.14985. Oncotarget. 2017. PMID: 28152522 Free PMC article.
-
VEGFC/VEGFR3 Signaling Regulates Mouse Spermatogonial Cell Proliferation via the Activation of AKT/MAPK and Cyclin D1 Pathway and Mediates the Apoptosis by affecting Caspase 3/9 and Bcl-2.Cell Cycle. 2018;17(2):225-239. doi: 10.1080/15384101.2017.1407891. Epub 2018 Jan 2. Cell Cycle. 2018. PMID: 29169284 Free PMC article.
-
Sperm-carried IGF2 downregulated the expression of mitogens produced by Sertoli cells: A paracrine mechanism for regulating spermatogenesis?Front Endocrinol (Lausanne). 2022 Nov 29;13:1010796. doi: 10.3389/fendo.2022.1010796. eCollection 2022. Front Endocrinol (Lausanne). 2022. PMID: 36523595 Free PMC article.
-
Nodal promotes the self-renewal of human colon cancer stem cells via an autocrine manner through Smad2/3 signaling pathway.Biomed Res Int. 2014;2014:364134. doi: 10.1155/2014/364134. Epub 2014 Feb 17. Biomed Res Int. 2014. PMID: 24696849 Free PMC article.
-
Sequential expression of long noncoding RNA as mRNA gene expression in specific stages of mouse spermatogenesis.Sci Rep. 2014 Aug 6;4:5966. doi: 10.1038/srep05966. Sci Rep. 2014. PMID: 25097017 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

