Molecular mechanisms by which human immunodeficiency virus type 1 integrase stimulates the early steps of reverse transcription
- PMID: 17626089
- PMCID: PMC2045400
- DOI: 10.1128/JVI.00519-07
Molecular mechanisms by which human immunodeficiency virus type 1 integrase stimulates the early steps of reverse transcription
Abstract
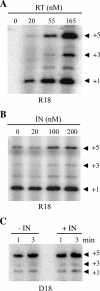
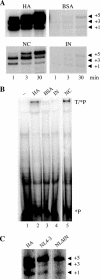
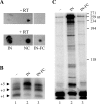
Reverse transcriptase (RT) and integrase (IN) are two essential enzymes that play a critical role in synthesis and integration of the retroviral cDNA, respectively. For human immunodeficiency virus type 1 (HIV-1), RT and IN physically interact and certain mutations and deletions of IN result in viruses defective in early steps of reverse transcription. However, the mechanism by which IN affects reverse transcription is not understood. We used a cell-free reverse transcription assay with different primers and compositions of deoxynucleoside triphosphates to differentially monitor the effect of IN on the initiation and elongation modes of reverse transcription. During the initiation mode, addition of IN stimulated RT-catalyzed reverse transcription by fourfold. The stimulation was specific to IN and could not be detected when the full-length IN was replaced with truncated IN derivatives. The IN-stimulated initiation was also restricted to the template-primer complex formed using tRNA(3)(Lys) or short RNA oligonucleotides as the primer and not those formed using DNA oligonucleotides as the primer. Addition of IN also produced a threefold stimulation during the elongation mode, which was not primer dependent. The stimulation of both initiation and elongation by IN was retained in the presence of an RT trap. Furthermore, IN had no effect on steps at or before template-primer annealing, including packaging of viral genomic RNA and tRNA(3)(Lys). Taken together, our results showed that IN acts at early steps of reverse transcription by increasing the processivity of RT and suppressing the formation of the pause products.
Figures




References
-
- Arts, E. J., M. Ghosh, P. S. Jacques, B. Ehresmann, and S. F. Le Grice. 1996. Restoration of tRNA3Lys-primed(-)-strand DNA synthesis to an HIV-1 reverse transcriptase mutant with extended tRNAs. Implications for retroviral replication. J. Biol. Chem. 271:9054-9061. - PubMed
-
- Arts, E. J., X. Li, Z. Gu, L. Kleiman, M. A. Parniak, and M. A. Wainberg. 1994. Comparison of deoxyoligonucleotide and tRNA(Lys-3) as primers in an endogenous human immunodeficiency virus-1 in vitro reverse transcription/template-switching reaction. J. Biol. Chem. 269:14672-14680. - PubMed
-
- Brown, P. O. 1997. Integration. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

