Monoubiquitylation of H2A.Z distinguishes its association with euchromatin or facultative heterochromatin
- PMID: 17636032
- PMCID: PMC2099601
- DOI: 10.1128/MCB.00241-07
Monoubiquitylation of H2A.Z distinguishes its association with euchromatin or facultative heterochromatin
Abstract
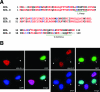
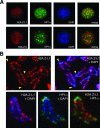
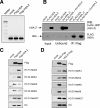
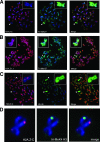
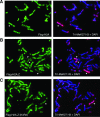
H2A.Z is a histone H2A variant that is essential for viability in organisms such as Tetrahymena thermophila, Drosophila melanogaster, and mice. In Saccharomyces cerevisiae, loss of H2A.Z is tolerated, but proper regulation of gene expression is affected. Genetics and genome-wide localization studies show that yeast H2A.Z physically localizes to the promoters of genes and functions in part to protect active genes in euchromatin from being silenced by heterochromatin spreading. To date, the function of H2A.Z in mammalian cells is less clear, and evidence so far suggests that it has a role in chromatin compaction and heterochromatin silencing. In this study, we found that the bulk of H2A.Z is excluded from constitutive heterochromatin in differentiated human and mouse cells. Consistent with this observation, analyses of H2A.Z- or H2A-containing mononucleosomes show that the H3 associated with H2A.Z has lower levels of K9 methylation but higher levels of K4 methylation than those associated with H2A. We also found that a fraction of mammalian H2A.Z is monoubiquitylated and that, on the inactive X chromosomes of female cells, the majority of this histone variant is modified by ubiquitin. Finally, ubiquitylation of H2A.Z is mediated by the RING1b E3 ligase of the human polycomb complex, further supporting a silencing role of ubiquitylated H2A.Z. These new findings suggest that mammalian H2A.Z is associated with both euchromatin and facultative heterochromatin and that monoubiquitylation is a specific mark that distinguishes the H2A.Z associated with these different chromatin states.
Figures



References
-
- Abbott, D. W., V. S. Ivanova, X. Wang, W. M. Bonner, and J. Ausio. 2001. Characterization of the stability and folding of H2A.Z chromatin particles: implications for transcriptional activation. J. Biol. Chem. 276:41945-41949. - PubMed
-
- Bannister, A. J., P. Zegerman, J. F. Partridge, E. A. Miska, J. O. Thomas, R. C. Allshire, and T. Kouzarides. 2001. Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 410:120-124. - PubMed
-
- Bernstein, E., and S. B. Hake. 2006. The nucleosome: a little variation goes a long way. Biochem. Cell Biol. 84:505-517. - PubMed
-
- Boggs, B. A., P. Cheung, E. Heard, D. L. Spector, A. C. Chinault, and C. D. Allis. 2002. Differentially methylated forms of histone H3 show unique association patterns with inactive human X chromosomes. Nat. Genet. 30:73-76. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
