Ancient and continuing Darwinian selection on insulin-like growth factor II in placental fishes
- PMID: 17636118
- PMCID: PMC1941482
- DOI: 10.1073/pnas.0705048104
Ancient and continuing Darwinian selection on insulin-like growth factor II in placental fishes
Abstract
Despite abundant examples of both adaptation at the level of phenotype and Darwinian selection at the level of genes, correlations between these two processes are notoriously difficult to identify. Positive Darwinian selection on genes is most easily discerned in cases of genetic conflict, when antagonistic evolutionary processes such as a Red Queen race drive the rate of nonsynonymous substitution above the neutral mutation rate. Genomic imprinting in mammals is thought to be the product of antagonistic evolution coincident with evolution of the placenta, but imprinted loci lack evidence of positive selection likely because of the ancient origin of viviparity in mammals. To determine whether genetic conflict is a general feature of adaptation to placental reproduction, we performed comparative evolutionary analyses of the insulin-like growth factor II (IGF2) gene in teleost fishes. Our analysis included several members of the order Cyprinodontiformes, in which livebearing and placentation have evolved several times independently. We found that IGF2 is subject to positive Darwinian selection coincident with the evolution of placentation in fishes, with particularly strong selection among lineages that have evolved placentation recently. Positive selection is also detected along ancient lineages of placental livebearing fishes, suggesting that selection on IGF2 function is ongoing in placental species. Our observations provide a rare example of natural selection acting in synchrony at the phenotypic and molecular level. These results also constitute the first direct evidence of parent-offspring conflict driving gene evolution.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
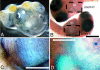
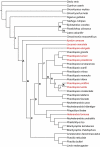


Similar articles
-
Sequence analyses of the distal-less homeobox gene family in East African cichlid fishes reveal signatures of positive selection.BMC Evol Biol. 2013 Jul 17;13:153. doi: 10.1186/1471-2148-13-153. BMC Evol Biol. 2013. PMID: 23865956 Free PMC article.
-
Positive Darwinian selection at the pantophysin (Pan I) locus in marine gadid fishes.Mol Biol Evol. 2004 Jan;21(1):65-75. doi: 10.1093/molbev/msg237. Epub 2003 Aug 29. Mol Biol Evol. 2004. PMID: 12949133
-
Positive Darwinian selection in the singularly large taste receptor gene family of an 'ancient' fish, Latimeria chalumnae.BMC Genomics. 2014 Aug 5;15(1):650. doi: 10.1186/1471-2164-15-650. BMC Genomics. 2014. PMID: 25091523 Free PMC article.
-
Genomic imprinting in marsupial placentation.Reproduction. 2008 Nov;136(5):523-31. doi: 10.1530/REP-08-0264. Epub 2008 Sep 19. Reproduction. 2008. PMID: 18805821 Review.
-
The origin and evolution of genomic imprinting and viviparity in mammals.Philos Trans R Soc Lond B Biol Sci. 2013 Jan 5;368(1609):20120151. doi: 10.1098/rstb.2012.0151. Philos Trans R Soc Lond B Biol Sci. 2013. PMID: 23166401 Free PMC article. Review.
Cited by
-
Functional similarity and molecular divergence of a novel reproductive transcriptome in two male-pregnant Syngnathus pipefish species.Ecol Evol. 2013 Oct;3(12):4092-108. doi: 10.1002/ece3.763. Epub 2013 Sep 20. Ecol Evol. 2013. PMID: 24324861 Free PMC article.
-
Retroviruses facilitate the rapid evolution of the mammalian placenta.Bioessays. 2013 Oct;35(10):853-61. doi: 10.1002/bies.201300059. Epub 2013 Jul 19. Bioessays. 2013. PMID: 23873343 Free PMC article.
-
Evidence of positive selection associated with placental loss in tiger sharks.BMC Evol Biol. 2016 Jun 14;16(1):126. doi: 10.1186/s12862-016-0696-y. BMC Evol Biol. 2016. PMID: 27296413 Free PMC article.
-
The genome of the live-bearing fish Heterandria formosa implicates a role of conserved vertebrate genes in the evolution of placental fish.BMC Evol Biol. 2019 Jul 26;19(1):156. doi: 10.1186/s12862-019-1484-2. BMC Evol Biol. 2019. PMID: 31349784 Free PMC article.
-
The evolution of the placenta drives a shift in sexual selection in livebearing fish.Nature. 2014 Sep 11;513(7517):233-6. doi: 10.1038/nature13451. Epub 2014 Jul 9. Nature. 2014. PMID: 25043015
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

