The multifunctional protein p54nrb/PSF recruits the exonuclease XRN2 to facilitate pre-mRNA 3' processing and transcription termination
- PMID: 17639083
- PMCID: PMC1920172
- DOI: 10.1101/gad.1565207
The multifunctional protein p54nrb/PSF recruits the exonuclease XRN2 to facilitate pre-mRNA 3' processing and transcription termination
Abstract
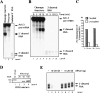
Termination of RNA polymerase II transcription frequently requires a poly(A) signal and cleavage/polyadenylation factors. Recent work has shown that degradation of the downstream cleaved RNA by the exonuclease XRN2 promotes termination, but how XRN2 functions with 3'-processing factors to elicit termination remains unclear. Here we show that XRN2 physically associates with 3'-processing factors and accumulates at the 3' end of a transcribed gene. In vitro 3'-processing assays show that XRN2 is necessary to degrade the downstream RNA, but is not required for 3' cleavage. Significantly, degradation of the 3'-cleaved RNA was stimulated when coupled to cleavage. Unexpectedly, while investigating how XRN2 is recruited to the 3'-processing machinery, we found that XRN2 associates with p54nrb/NonO(p54)-protein-associated splicing factor (PSF), multifunctional proteins involved in several nuclear processes. Strikingly, p54 is also required for degradation of the 3'-cleaved RNA in vitro. p54 is present along the length of genes, and small interfering RNA (siRNA)-mediated knockdown leads to defects in XRN2 recruitment and termination. Together, our data indicate that p54nrb/PSF functions in recruitment of XRN2 to facilitate pre-mRNA 3' processing and transcription termination.
Figures
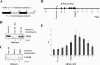





Similar articles
-
Splicing and transcription-associated proteins PSF and p54nrb/nonO bind to the RNA polymerase II CTD.RNA. 2002 Sep;8(9):1102-11. doi: 10.1017/s1355838202025037. RNA. 2002. PMID: 12358429 Free PMC article.
-
p54nrb is a new regulator of progression of malignant melanoma.Carcinogenesis. 2011 Aug;32(8):1176-82. doi: 10.1093/carcin/bgr103. Epub 2011 Jun 3. Carcinogenesis. 2011. PMID: 21642354
-
p54nrb/NONO regulates cyclic AMP-dependent glucocorticoid production by modulating phosphodiesterase mRNA splicing and degradation.Mol Cell Biol. 2015 Apr;35(7):1223-37. doi: 10.1128/MCB.00993-14. Epub 2015 Jan 20. Mol Cell Biol. 2015. PMID: 25605330 Free PMC article.
-
Paraspeckles: nuclear bodies built on long noncoding RNA.J Cell Biol. 2009 Sep 7;186(5):637-44. doi: 10.1083/jcb.200906113. Epub 2009 Aug 31. J Cell Biol. 2009. PMID: 19720872 Free PMC article. Review.
-
PSF and p54(nrb)/NonO--multi-functional nuclear proteins.FEBS Lett. 2002 Nov 6;531(2):109-14. doi: 10.1016/s0014-5793(02)03447-6. FEBS Lett. 2002. PMID: 12417296 Review.
Cited by
-
R Loops and Links to Human Disease.J Mol Biol. 2017 Oct 27;429(21):3168-3180. doi: 10.1016/j.jmb.2016.08.031. Epub 2016 Sep 4. J Mol Biol. 2017. PMID: 27600412 Free PMC article. Review.
-
Characterization of DNA binding and pairing activities associated with the native SFPQ·NONO DNA repair protein complex.Biochem Biophys Res Commun. 2015 Aug 7;463(4):473-8. doi: 10.1016/j.bbrc.2015.05.024. Epub 2015 May 18. Biochem Biophys Res Commun. 2015. PMID: 25998385 Free PMC article.
-
Established and Evolving Roles of the Multifunctional Non-POU Domain-Containing Octamer-Binding Protein (NonO) and Splicing Factor Proline- and Glutamine-Rich (SFPQ).J Dev Biol. 2024 Jan 5;12(1):3. doi: 10.3390/jdb12010003. J Dev Biol. 2024. PMID: 38248868 Free PMC article. Review.
-
Fox-3 and PSF interact to activate neural cell-specific alternative splicing.Nucleic Acids Res. 2011 Apr;39(8):3064-78. doi: 10.1093/nar/gkq1221. Epub 2010 Dec 21. Nucleic Acids Res. 2011. PMID: 21177649 Free PMC article.
-
The RNA-binding protein SFPQ preserves long-intron splicing and regulates circRNA biogenesis in mammals.Elife. 2021 Jan 21;10:e63088. doi: 10.7554/eLife.63088. Elife. 2021. PMID: 33476259 Free PMC article.
References
-
- Ahn S.H., Kim M., Buratowski S., Kim M., Buratowski S., Buratowski S. Phosphorylation of serine 2 within the RNA polymerase II C-terminal domain couples transcription and 3′ end processing. Mol. Cell. 2004;13:67–76. - PubMed
-
- Bashkirov V.I., Scherthan H., Solinger J.A., Buerstedde J.M., Heyer W.D., Scherthan H., Solinger J.A., Buerstedde J.M., Heyer W.D., Solinger J.A., Buerstedde J.M., Heyer W.D., Buerstedde J.M., Heyer W.D., Heyer W.D. A mouse cytoplasmic exoribonuclease (mXRN1p) with preference for G4 tetraplex substrates. J. Cell Biol. 1997;136:761–773. - PMC - PubMed
-
- Birse C.E., Minvielle-Sebastia L., Lee B.A., Keller W., Proudfoot N.J., Minvielle-Sebastia L., Lee B.A., Keller W., Proudfoot N.J., Lee B.A., Keller W., Proudfoot N.J., Keller W., Proudfoot N.J., Proudfoot N.J. Coupling termination of transcription to messenger RNA maturation in yeast. Science. 1998;280:298–301. - PubMed
-
- Bousquet-Antonelli C., Presutti C., Tollervey D., Presutti C., Tollervey D., Tollervey D. Identification of a regulated pathway for nuclear pre-mRNA turnover. Cell. 2000;102:765–775. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
