Co-localization of CENP-C and CENP-H to discontinuous domains of CENP-A chromatin at human neocentromeres
- PMID: 17651496
- PMCID: PMC2323242
- DOI: 10.1186/gb-2007-8-7-r148
Co-localization of CENP-C and CENP-H to discontinuous domains of CENP-A chromatin at human neocentromeres
Abstract
Background: Mammalian centromere formation is dependent on chromatin that contains centromere protein (CENP)-A, which is the centromere-specific histone H3 variant. Human neocentromeres have acquired CENP-A chromatin epigenetically in ectopic chromosomal locations on low-copy complex DNA. Neocentromeres permit detailed investigation of centromeric chromatin organization that is not possible in the highly repetitive alpha satellite DNA present at endogenous centromeres.
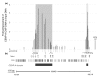
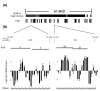
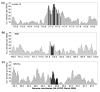
Results: We have examined the distribution of CENP-A, as well as two additional centromeric chromatin-associated proteins (CENP-C and CENP-H), across neocentromeric DNA using chromatin immunoprecipitation (ChIP) on CHIP assays on custom genomic microarrays at three different resolutions. Analysis of two neocentromeres using a contiguous bacterial artificial chromosome (BAC) microarray spanning bands 13q31.3 to 13q33.1 shows that both CENP-C and CENP-H co-localize to the CENP-A chromatin domain. Using a higher resolution polymerase chain reaction (PCR)-amplicon microarray spanning the neocentromere, we find that the CENP-A chromatin is discontinuous, consisting of a major domain of about 87.8 kilobases (kb) and a minor domain of about 13.2 kb, separated by an approximately 158 kb region devoid of CENPs. Both CENP-A domains exhibit co-localization of CENP-C and CENP-H, defining a distinct inner kinetochore chromatin structure that is consistent with higher order chromatin looping models at centromeres. The PCR microarray data suggested varying density of CENP-A nucleosomes across the major domain, which was confirmed using a higher resolution oligo-based microarray.
Conclusion: Centromeric chromatin consists of several CENP-A subdomains with highly discontinuous CENP-A chromatin at both the level of individual nucleosomes and at higher order chromatin levels, raising questions regarding the overall structure of centromeric chromatin.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

