Physical-chemical determinants of turn conformations in globular proteins
- PMID: 17656584
- PMCID: PMC2203374
- DOI: 10.1110/ps.072898507
Physical-chemical determinants of turn conformations in globular proteins
Abstract
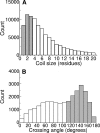
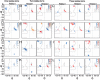
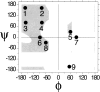
Globular proteins are assemblies of alpha-helices and beta-strands, interconnected by reverse turns and longer loops. Most short turns can be classified readily into a limited repertoire of discrete backbone conformations, but the physical-chemical determinants of these distinct conformational basins remain an open question. We investigated this question by exhaustive analysis of all backbone conformations accessible to short chain segments bracketed by either an alpha-helix or a beta-strand (i.e., alpha-segment-alpha, beta-segment-beta, alpha-segment-beta, and beta-segment-alpha) in a nine-state model. We find that each of these four secondary structure environments imposes its own unique steric and hydrogen-bonding constraints on the intervening segment, resulting in a limited repertoire of conformations. In greater detail, an exhaustive set of conformations was generated for short backbone segments having reverse-turn chain topology and bracketed between elements of secondary structure. This set was filtered, and only clash-free, hydrogen-bond-satisfied conformers having reverse-turn topology were retained. The filtered set includes authentic turn conformations, observed in proteins of known structure, but little else. In particular, over 99% of the alternative conformations failed to satisfy at least one criterion and were excluded from the filtered set. Furthermore, almost all of the remaining alternative conformations have close tolerances that would be too tight to accommodate side chains longer than a single beta-carbon. These results provide a molecular explanation for the observation that reverse turns between elements of regular secondary can be classified into a small number of discrete conformations.
Figures


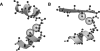
Similar articles
-
Expanded turn conformations: characterization and sequence-structure correspondence in alpha-turns with implications in helix folding.Proteins. 2004 May 1;55(2):305-15. doi: 10.1002/prot.20064. Proteins. 2004. PMID: 15048823
-
Understanding the role of the topology in protein folding by computational inverse folding experiments.Comput Biol Chem. 2008 Aug;32(4):233-9. doi: 10.1016/j.compbiolchem.2008.03.015. Epub 2008 Apr 8. Comput Biol Chem. 2008. PMID: 18479970
-
Characterization of beta-turn and Asx-turns mimicry in a model peptide: stabilization via C--H . . . O interaction.Biopolymers. 2006 Apr 15;81(6):440-9. doi: 10.1002/bip.20441. Biopolymers. 2006. PMID: 16411188
-
Prediction of tight turns and their types in proteins.Anal Biochem. 2000 Nov 1;286(1):1-16. doi: 10.1006/abio.2000.4757. Anal Biochem. 2000. PMID: 11038267 Review.
-
Helices in peptoids of alpha- and beta-peptides.Phys Biol. 2006 Feb 2;3(1):S1-9. doi: 10.1088/1478-3975/3/1/S01. Phys Biol. 2006. PMID: 16582460 Review.
Cited by
-
Principles for designing ideal protein structures.Nature. 2012 Nov 8;491(7423):222-7. doi: 10.1038/nature11600. Nature. 2012. PMID: 23135467 Free PMC article.
-
Stabilization of the spectrin-like domains of nesprin-1α by the evolutionarily conserved "adaptive" domain.Cell Mol Bioeng. 2010 Jun 1;3(2):139-150. doi: 10.1007/s12195-010-0121-3. Cell Mol Bioeng. 2010. PMID: 20563238 Free PMC article.
-
The importance of slow motions for protein functional loops.Phys Biol. 2012 Feb;9(1):014001. doi: 10.1088/1478-3975/9/1/014001. Epub 2012 Feb 7. Phys Biol. 2012. PMID: 22314977 Free PMC article.
-
Reducing the dimensionality of the protein-folding search problem.Protein Sci. 2012 Aug;21(8):1231-40. doi: 10.1002/pro.2106. Epub 2012 Jul 6. Protein Sci. 2012. PMID: 22692765 Free PMC article.
-
On the occurrence of linear groups in proteins.Protein Sci. 2009 Jun;18(6):1321-5. doi: 10.1002/pro.133. Protein Sci. 2009. PMID: 19472372 Free PMC article.
References
-
- Baldwin R.L. 2003. In search of the energetic role of peptide hydrogen bonds. J. Biol. Chem. 278: 17581–17588. - PubMed
-
- Brunet A.P., Huang, E.S., Huffine, M.E., Loeb, J.E., Weltman, R.J., and Hecht, M.H. 1993. The role of turns in the structure of an α-helical protein. Nature 364: 355–358. - PubMed
-
- Chothia C., Levitt, M., and Richardson, D. 1981. Helix to helix packing in proteins. J. Mol. Biol. 145: 215–250. - PubMed
-
- Chou P.Y. and Fasman, G.D. 1977. β-Turns in proteins. J. Mol. Biol. 115: 135–175. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

