Novel tonoplast transporters identified using a proteomic approach with vacuoles isolated from cauliflower buds
- PMID: 17660356
- PMCID: PMC1976570
- DOI: 10.1104/pp.107.096917
Novel tonoplast transporters identified using a proteomic approach with vacuoles isolated from cauliflower buds
Abstract
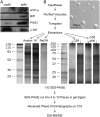
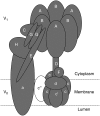
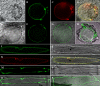
Young meristematic plant cells contain a large number of small vacuoles, while the largest part of the vacuome in mature cells is composed by a large central vacuole, occupying 80% to 90% of the cell volume. Thus far, only a limited number of vacuolar membrane proteins have been identified and characterized. The proteomic approach is a powerful tool to identify new vacuolar membrane proteins. To analyze vacuoles from growing tissues we isolated vacuoles from cauliflower (Brassica oleracea) buds, which are constituted by a large amount of small cells but also contain cells in expansion as well as fully expanded cells. Here we show that using purified cauliflower vacuoles and different extraction procedures such as saline, NaOH, acetone, and chloroform/methanol and analyzing the data against the Arabidopsis (Arabidopsis thaliana) database 102 cauliflower integral proteins and 214 peripheral proteins could be identified. The vacuolar pyrophosphatase was the most prominent protein. From the 102 identified proteins 45 proteins were already described. Nine of these, corresponding to 46% of peptides detected, are known vacuolar proteins. We identified 57 proteins (55.9%) containing at least one membrane spanning domain with unknown subcellular localization. A comparison of the newly identified proteins with expression profiles from in silico data revealed that most of them are highly expressed in young, developing tissues. To verify whether the newly identified proteins were indeed localized in the vacuole we constructed and expressed green fluorescence protein fusion proteins for five putative vacuolar membrane proteins exhibiting three to 11 transmembrane domains. Four of them, a putative organic cation transporter, a nodulin N21 family protein, a membrane protein of unknown function, and a senescence related membrane protein were localized in the vacuolar membrane, while a white-brown ATP-binding cassette transporter homolog was shown to reside in the plasma membrane. These results demonstrate that proteomic analysis of highly purified vacuoles from specific tissues allows the identification of new vacuolar proteins and provides an additional view of tonoplastic proteins.
Figures


References
-
- Apse MP, Aharon GS, Snedden WA, Blumwald E (1999) Salt tolerance conferred by overexpression of a vacuolar Na+/H+ antiport in Arabidopsis. Science 285 1256–1258 - PubMed
-
- Barrieu F, Thomas D, Marty-Mazars D, Charbonnier M, Marty F (1998) Tonoplast intrinsic proteins from cauliflower (Brassica oleracea L. var. botrytis): immunological analysis, cDNA cloning and evidence for expression in meristematic tissues. Planta 204 335–344 - PubMed
-
- Bednarczyk D, Ekins S, Wikel JH, Wright SH (2003) Influence of molecular structure on substrate binding to the human organic cation transporter, hOCT1. Mol Pharmacol 63 489–498 - PubMed
-
- Bradford MM (1976) Rapid and quantitative method for quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72 248–252 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

