EGFR associated expression profiles vary with breast tumor subtype
- PMID: 17663798
- PMCID: PMC2014778
- DOI: 10.1186/1471-2164-8-258
EGFR associated expression profiles vary with breast tumor subtype
Abstract
Background: The epidermal growth factor receptor (EGFR/HER1) and its downstream signaling events are important for regulating cell growth and behavior in many epithelial tumors types. In breast cancer, the role of EGFR is complex and appears to vary relative to important clinical features including estrogen receptor (ER) status. To investigate EGFR-signaling using a genomics approach, several breast basal-like and luminal epithelial cell lines were examined for sensitivity to EGFR inhibitors. An EGFR-associated gene expression signature was identified in the basal-like SUM102 cell line and was used to classify a diverse set of sporadic breast tumors.
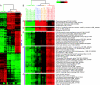
Results: In vitro, breast basal-like cell lines were more sensitive to EGFR inhibitors compared to luminal cell lines. The basal-like tumor derived lines were also the most sensitive to carboplatin, which acted synergistically with cetuximab. An EGFR-associated signature was developed in vitro, evaluated on 241 primary breast tumors; three distinct clusters of genes were evident in vivo, two of which were predictive of poor patient outcomes. These EGFR-associated poor prognostic signatures were highly expressed in almost all basal-like tumors and many of the HER2+/ER- and Luminal B tumors.
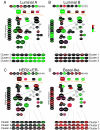
Conclusion: These results suggest that breast basal-like cell lines are sensitive to EGFR inhibitors and carboplatin, and this combination may also be synergistic. In vivo, the EGFR-signatures were of prognostic value, were associated with tumor subtype, and were uniquely associated with the high expression of distinct EGFR-RAS-MEK pathway genes.
Figures



References
-
- Sørlie T, Perou CM, Tibshirani R, Aas T, Geisler S, Johnsen H, Hastie T, Eisen MB, van de Rijn M, Jeffrey SS, Thorsen T, Quist H, Matese JC, Brown PO, Botstein D, Eystein Lonning P, Borresen-Dale AL. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci U S A. 2001;98:10869–10874. doi: 10.1073/pnas.191367098. - DOI - PMC - PubMed
-
- Sørlie T, Tibshirani R, Parker J, Hastie T, Marron JS, Nobel A, Deng S, Johnsen H, Pesich R, Geisler S, Demeter J, Perou CM, Lonning PE, Brown PO, Borresen-Dale AL, Botstein D. Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc Natl Acad Sci U S A. 2003;100:8418–8423. doi: 10.1073/pnas.0932692100. - DOI - PMC - PubMed
-
- Perou CM, Sørlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, Pollack JR, Ross DT, Johnsen H, Akslen LA, Fluge O, Pergamenschikov A, Williams C, Zhu SX, Lonning PE, Borresen-Dale AL, Brown PO, Botstein D. Molecular portraits of human breast tumours. Nature. 2000;406:747–752. doi: 10.1038/35021093. - DOI - PubMed
-
- Hu Z, Troester M, Perou CM. High reproducibility using sodium hydroxide-stripped long oligonucleotide DNA microarrays. Biotechniques. 2005;38:121–124. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

