The structure of a putative scaffolding protein of immature poxvirus particles as determined by electron microscopy suggests similarity with capsid proteins of large icosahedral DNA viruses
- PMID: 17670837
- PMCID: PMC2045580
- DOI: 10.1128/JVI.00594-07
The structure of a putative scaffolding protein of immature poxvirus particles as determined by electron microscopy suggests similarity with capsid proteins of large icosahedral DNA viruses
Abstract
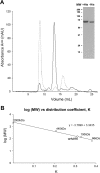
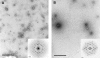
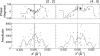
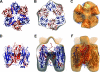
Orf virus, the prototype parapoxvirus, is responsible for contagious ecthyma in sheep and goats. The central region of the viral genome codes for proteins highly conserved among vertebrate poxviruses and which are frequently essential for viral proliferation. Analysis of the recently published genome sequence of orf virus revealed that among such essential proteins, the protein orfv075 is an orthologue of D13, the rifampin resistance gene product critical for vaccinia virus morphogenesis. Previous studies showed that D13, arranged as "spicules," is necessary for the formation of vaccinia virus immature virions, a mandatory intermediate in viral maturation. We have determined the three-dimensional structure of recombinant orfv075 at approximately 25-A resolution by electron microscopy of two-dimensional crystals. orfv075 organizes as trimers with a tripod-like main body and a propeller-like smaller domain. The molecular envelope of orfv075 shows unexpectedly good agreement to that of a distant homologue, VP54, the major capsid protein of Paramecium bursaria Chlorella virus type 1. Our structural analysis suggests that orfv075 belongs in the double-barreled capsid protein family found in many double-stranded DNA icosahedral viruses and supports the hypothesis that the nonicosahedral poxviruses and the large icosahedral DNA viruses are evolutionarily related.
Figures




Similar articles
-
Cryo-EM structure of orf virus scaffolding protein orfv075.Biochem Biophys Res Commun. 2024 Oct 8;728:150334. doi: 10.1016/j.bbrc.2024.150334. Epub 2024 Jun 29. Biochem Biophys Res Commun. 2024. PMID: 38968773
-
External scaffold of spherical immature poxvirus particles is made of protein trimers, forming a honeycomb lattice.J Cell Biol. 2005 Sep 12;170(6):971-81. doi: 10.1083/jcb.200504026. Epub 2005 Sep 6. J Cell Biol. 2005. PMID: 16144903 Free PMC article.
-
The capsid proteins of a large, icosahedral dsDNA virus.J Mol Biol. 2009 Jan 30;385(4):1287-99. doi: 10.1016/j.jmb.2008.11.002. Epub 2008 Nov 12. J Mol Biol. 2009. PMID: 19027752 Free PMC article.
-
Structures of giant icosahedral eukaryotic dsDNA viruses.Curr Opin Virol. 2011 Aug;1(2):101-9. doi: 10.1016/j.coviro.2011.06.005. Curr Opin Virol. 2011. PMID: 21909343 Free PMC article. Review.
-
In a nutshell: structure and assembly of the vaccinia virion.Adv Virus Res. 2006;66:31-124. doi: 10.1016/S0065-3527(06)66002-8. Adv Virus Res. 2006. PMID: 16877059 Review.
Cited by
-
Poxvirus under the eyes of electron microscope.Appl Microsc. 2022 Nov 14;52(1):11. doi: 10.1186/s42649-022-00080-3. Appl Microsc. 2022. PMID: 36372822 Free PMC article. Review.
-
Deletion of the Chemokine Binding Protein Gene from the Parapoxvirus Orf Virus Reduces Virulence and Pathogenesis in Sheep.Front Microbiol. 2017 Jan 24;8:46. doi: 10.3389/fmicb.2017.00046. eCollection 2017. Front Microbiol. 2017. PMID: 28174562 Free PMC article.
-
Oligomeric structure of colicin ia channel in lipid bilayer membranes.J Biol Chem. 2009 Jun 12;284(24):16126-16134. doi: 10.1074/jbc.M900292200. Epub 2009 Apr 8. J Biol Chem. 2009. PMID: 19357078 Free PMC article.
-
Pass the jelly rolls.Structure. 2011 Jul 13;19(7):904-6. doi: 10.1016/j.str.2011.06.004. Structure. 2011. PMID: 21742257 Free PMC article.
-
Poxvirus membrane biogenesis.Virology. 2015 May;479-480:619-26. doi: 10.1016/j.virol.2015.02.003. Epub 2015 Feb 26. Virology. 2015. PMID: 25728299 Free PMC article. Review.
References
-
- Abrescia, N. G., J. J. Cockburn, J. M. Grimes, G. C. Sutton, J. M. Diprose, S. J. Butcher, S. D. Fuller, C. San Martin, R. M. Burnett, D. I. Stuart, D. H. Bamford, and J. K. Bamford. 2004. Insights into assembly from structural analysis of bacteriophage PRD1. Nature 432:68-74. - PubMed
-
- Amos, L. A., R. Henderson, and P. N. Unwin. 1982. Three-dimensional structure determination by electron microscopy of two-dimensional crystals. Prog. Biophys. Mol. Biol. 39:183-231. - PubMed
-
- Andrews, P. 1962. Estimation of molecular weights of proteins by gel filtration. Nature 196:36-39. - PubMed
-
- Baldick, C. J., Jr., and B. Moss. 1987. Resistance of vaccinia virus to rifampicin conferred by a single nucleotide substitution near the predicted NH2 terminus of a gene encoding an Mr 62,000 polypeptide. Virology 156:138-145. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

