Adaptive evolution has targeted the C-terminal domain of the RXLR effectors of plant pathogenic oomycetes
- PMID: 17675403
- PMCID: PMC2002621
- DOI: 10.1105/tpc.107.051037
Adaptive evolution has targeted the C-terminal domain of the RXLR effectors of plant pathogenic oomycetes
Abstract
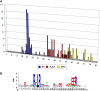
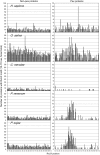
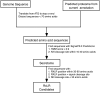
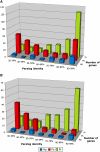
Oomycete plant pathogens deliver effector proteins inside host cells to modulate plant defense circuitry and to enable parasitic colonization. These effectors are defined by a conserved motif, termed RXLR (for Arg, any amino acid, Leu, Arg), that is located downstream of the signal peptide and that has been implicated in host translocation. Because the phenotypes of RXLR effectors extend to plant cells, their genes are expected to be the direct target of the evolutionary forces that drive the antagonistic interplay between pathogen and host. We used the draft genome sequences of three oomycete plant pathogens, Phytophthora sojae, Phytophthora ramorum, and Hyaloperonospora parasitica, to generate genome-wide catalogs of RXLR effector genes and determine the extent to which these genes are under positive selection. These analyses revealed that the RXLR sequence is overrepresented and positionally constrained in the secretome of Phytophthora relative to other eukaryotes. The three examined plant pathogenic oomycetes carry complex and diverse sets of RXLR effector genes that have undergone relatively rapid birth and death evolution. We obtained robust evidence of positive selection in more than two-thirds of the examined paralog families of RXLR effectors. Positive selection has acted for the most part on the C-terminal region, consistent with the view that RXLR effectors are modular, with the N terminus involved in secretion and host translocation and the C-terminal domain dedicated to modulating host defenses inside plant cells.
Figures

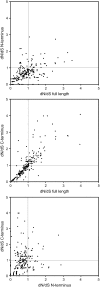



Similar articles
-
Homologous RXLR effectors from Hyaloperonospora arabidopsidis and Phytophthora sojae suppress immunity in distantly related plants.Plant J. 2012 Dec;72(6):882-93. doi: 10.1111/j.1365-313X.2012.05079.x. Epub 2012 Oct 26. Plant J. 2012. PMID: 22709376
-
Plasmodium falciparum and Hyaloperonospora parasitica effector translocation motifs are functional in Phytophthora infestans.Microbiology (Reading). 2008 Dec;154(Pt 12):3743-3751. doi: 10.1099/mic.0.2008/021964-0. Microbiology (Reading). 2008. PMID: 19047742
-
Towards understanding the virulence functions of RXLR effectors of the oomycete plant pathogen Phytophthora infestans.J Exp Bot. 2009;60(4):1133-40. doi: 10.1093/jxb/ern353. Epub 2009 Feb 9. J Exp Bot. 2009. PMID: 19204033 Review.
-
Short Linear Motifs (SLiMs) in "Core" RxLR Effectors of Phytophthora parasitica var. nicotianae: a Case of PpRxLR1 Effector.Microbiol Spectr. 2022 Apr 27;10(2):e0177421. doi: 10.1128/spectrum.01774-21. Epub 2022 Apr 11. Microbiol Spectr. 2022. PMID: 35404090 Free PMC article.
-
Oomycete RXLR effectors: delivery, functional redundancy and durable disease resistance.Curr Opin Plant Biol. 2008 Aug;11(4):373-9. doi: 10.1016/j.pbi.2008.04.005. Epub 2008 May 27. Curr Opin Plant Biol. 2008. PMID: 18511334 Review.
Cited by
-
The pathogenic mechanisms of Tilletia horrida as revealed by comparative and functional genomics.Sci Rep. 2018 Oct 18;8(1):15413. doi: 10.1038/s41598-018-33752-w. Sci Rep. 2018. PMID: 30337609 Free PMC article.
-
Arms race: diverse effector proteins with conserved motifs.Plant Signal Behav. 2019;14(2):1557008. doi: 10.1080/15592324.2018.1557008. Epub 2019 Jan 9. Plant Signal Behav. 2019. PMID: 30621489 Free PMC article.
-
Gene duplication and fragment recombination drive functional diversification of a superfamily of cytoplasmic effectors in Phytophthora sojae.PLoS One. 2013 Jul 29;8(7):e70036. doi: 10.1371/journal.pone.0070036. Print 2013. PLoS One. 2013. PMID: 23922898 Free PMC article.
-
Pathogen virulence of Phytophthora infestans: from gene to functional genomics.Physiol Mol Biol Plants. 2013 Apr;19(2):165-77. doi: 10.1007/s12298-012-0157-z. Physiol Mol Biol Plants. 2013. Retraction in: Physiol Mol Biol Plants. 2015 Jan;21(1):167. doi: 10.1007/s12298-014-0263-1. PMID: 24431484 Free PMC article. Retracted. Review.
-
Phosphite protects Fagus sylvatica seedlings towards Phytophthora plurivora via local toxicity, priming and facilitation of pathogen recognition.PLoS One. 2014 Jan 28;9(1):e87860. doi: 10.1371/journal.pone.0087860. eCollection 2014. PLoS One. 2014. PMID: 24489973 Free PMC article.
References
-
- Allen, R.L., Bittner-Eddy, P.D., Grenville-Briggs, L.J., Meitz, J.C., Rehmany, A.P., Rose, L.E., and Beynon, J.L. (2004). Host-parasite coevolutionary conflict between Arabidopsis and downy mildew. Science 306 1957–1960. - PubMed
-
- Anisimova, M., Bielawski, J.P., and Yang, Z. (2001). Accuracy and power of the likelihood ratio test in detecting adaptive molecular evolution. Mol. Biol. Evol. 18 1585–1592. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

