The MicroRNA miR-124 promotes neuronal differentiation by triggering brain-specific alternative pre-mRNA splicing
- PMID: 17679093
- PMCID: PMC3139456
- DOI: 10.1016/j.molcel.2007.07.015
The MicroRNA miR-124 promotes neuronal differentiation by triggering brain-specific alternative pre-mRNA splicing
Abstract
Both microRNAs and alternative pre-mRNA splicing have been implicated in the development of the nervous system (NS), but functional interactions between these two pathways are poorly understood. We demonstrate that the neuron-specific microRNA miR-124 directly targets PTBP1 (PTB/hnRNP I) mRNA, which encodes a global repressor of alternative pre-mRNA splicing in nonneuronal cells. Among the targets of PTBP1 is a critical cassette exon in the pre-mRNA of PTBP2 (nPTB/brPTB/PTBLP), an NS-enriched PTBP1 homolog. When this exon is skipped, PTBP2 mRNA is subject to nonsense-mediated decay (NMD). During neuronal differentiation, miR-124 reduces PTBP1 levels, leading to the accumulation of correctly spliced PTBP2 mRNA and a dramatic increase in PTBP2 protein. These events culminate in the transition from non-NS to NS-specific alternative splicing patterns. We also present evidence that miR-124 plays a key role in the differentiation of progenitor cells to mature neurons. Thus, miR-124 promotes NS development, at least in part by regulating an intricate network of NS-specific alternative splicing.
Figures

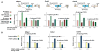





References
-
- Ashraf SI, McLoon AL, Sclarsic SM, Kunes S. Synaptic protein synthesis associated with memory is regulated by the RISC pathway in Drosophila. Cell. 2006;124:191–205. - PubMed
-
- Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–297. - PubMed
-
- Black DL. Mechanisms of alternative pre-messenger RNA splicing. Annu Rev Biochem. 2003;72:291–336. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

