Automated identification of analyzable metaphase chromosomes depicted on microscopic digital images
- PMID: 17681496
- PMCID: PMC2440955
- DOI: 10.1016/j.jbi.2007.06.008
Automated identification of analyzable metaphase chromosomes depicted on microscopic digital images
Abstract
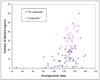
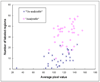
Visual search and identification of analyzable metaphase chromosomes using optical microscopes is a very tedious and time-consuming task that is routinely performed in genetic laboratories to detect and diagnose cancers and genetic diseases. The purpose of this study is to develop and test a computerized scheme that can automatically identify chromosomes in metaphase stage and classify them into analyzable and un-analyzable groups. Two independent datasets involving 170 images are used to train and test the scheme. The scheme uses image filtering, threshold, and labeling algorithms to detect chromosomes, followed by computing a set of features for each individual chromosome as well as for each identified metaphase cell. Two machine learning classifiers including a decision tree (DT) based on the features of individual chromosomes and an artificial neural network (ANN) using the features of the metaphase cells are optimized and tested to classify between analyzable and un-analyzable cells. Using the DT based classifier the Kappa coefficients for agreement between the cytogeneticist and the scheme are 0.83 and 0.89 for the training and testing datasets, respectively. We apply an independent testing and a 2-fold cross-validation method to assess the performance of the ANN-based classifier. The area under and receiver operating characteristic (ROC) curve is 0.93 for the complete dataset. This preliminary study demonstrates the feasibility of developing a computerized scheme to automatically identify and classify metaphase chromosomes.
Figures




References
-
- Tjio JH, Levan A. The chromosome number in man. Hereditas. 1956;42:1–6.
-
- Nowell PC, Hungerford DA. A minute chromosome in human chronic granulocytic leukemia. Science. 1960;132:1497. - PubMed
-
- Piper J, Granum E, Rutovitz D, Ruttledge H. Automation of chromosome analysis. Signal Processing. 1980;2:203–221.
-
- Truong K, Gibaud A, Zalcman G, Guilly M. Quantitative fish determination of chromosome 3 arm imbalances in lung tumors by automated image cytometry. Med Sci Monit. 2004;10:426–432. - PubMed
-
- Shih LM, Wang TL. Apply innovative technologies to explore cancer genome. Curr Opin Oncol. 2005;17:33–38. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

