An exfoliation and enrichment strategy results in improved transcriptional profiles when compared to matched formalin fixed samples
- PMID: 17683544
- PMCID: PMC1955442
- DOI: 10.1186/1472-6890-7-7
An exfoliation and enrichment strategy results in improved transcriptional profiles when compared to matched formalin fixed samples
Abstract
Background: Identifying the influence formalin fixation has on RNA integrity and recovery from clinical tissue specimens is integral to determining the utility of using archival tissue blocks in future molecular studies. For clinical material, the current gold standard is unfixed tissue that has been snap frozen. Fixed and frozen tissue however, both require laser capture microdissection to select for a specific cell population to study. The recent development of a sampling method capable of obtaining a viable, enriched cell population represents an alternative option in procuring cells from clinical material for molecular research purposes. The expression profiles of cells obtained by using this procurement approach, in conjunction with the profiles from cells laser capture microdissected from frozen tissue sections, were compared to the expression profiles from formalin fixed cells to determine the influence fixation has on expression profiles in clinical material.
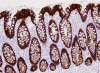
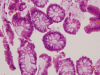
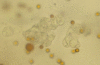
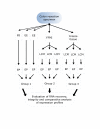
Methods: Triplicate samples of non-neoplastic colonic epithelial cells were recovered from a hemicolectomy specimen using three different procurement methods from the same originating site: 1) an exfoliation and enrichment strategy 2) laser capture microdissection from formalin fixed tissue and 3) laser capture microdissection from frozen tissue. Parameters currently in use to assess RNA integrity were utilized to assess the quality of recovered RNA. Additionally, an expression microarray was performed on each sample to assess the influence each procurement technique had on RNA recovery and degradation.
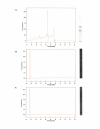
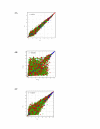
Results: The exfoliation/enrichment strategy was quantitatively and qualitatively superior to tissue that was formalin fixed. Fixation negatively influenced the expression profile of the formalin fixed group compared to both the frozen and exfoliated/enrichment groups.
Conclusion: The exfoliation/enrichment technique represents a superior alternative in tissue procurement and RNA recovery relative to formalin fixed tissue. None of the deleterious effects associated with formalin fixation are encountered in the exfoliated/enriched samples because of the absence of its use in this protocol. The exfoliation/enrichment technique also represents an economical alternative that will yield comparable results to cells enriched by laser capture microdissection from frozen tissue sections.
Figures

Similar articles
-
Manual exfoliation of fresh tissue obviates the need for frozen sections for molecular profiling.Cancer. 2005 Dec 25;105(6):483-91. doi: 10.1002/cncr.21347. Cancer. 2005. PMID: 16015639
-
Quantitative gene expression analysis in microdissected archival formalin-fixed and paraffin-embedded tumor tissue.Am J Pathol. 2001 Feb;158(2):419-29. doi: 10.1016/S0002-9440(10)63985-5. Am J Pathol. 2001. PMID: 11159180 Free PMC article.
-
Gene expression analysis in microdissected samples from decalcified tissues.Diagn Mol Pathol. 2012 Jun;21(2):120-6. doi: 10.1097/PDM.0b013e31823e9395. Diagn Mol Pathol. 2012. PMID: 22555095
-
Complementary techniques: laser capture microdissection--increasing specificity of gene expression profiling of cancer specimens.Adv Exp Med Biol. 2007;593:54-65. doi: 10.1007/978-0-387-39978-2_6. Adv Exp Med Biol. 2007. PMID: 17265716 Review.
-
Tissue microdissection.Methods Mol Biol. 2008;424:433-48. doi: 10.1007/978-1-60327-064-9_34. Methods Mol Biol. 2008. PMID: 18369881 Review.
Cited by
-
Survival of exfoliated epithelial cells: a delicate balance between anoikis and apoptosis.J Biomed Biotechnol. 2011;2011:534139. doi: 10.1155/2011/534139. Epub 2011 Oct 27. J Biomed Biotechnol. 2011. PMID: 22131811 Free PMC article. Review.
-
Clinical implementation of RNA signatures for pharmacogenomic decision-making.Pharmgenomics Pers Med. 2011;4:95-107. doi: 10.2147/PGPM.S14888. Epub 2011 Sep 8. Pharmgenomics Pers Med. 2011. PMID: 23226056 Free PMC article.
-
Biospecimen reporting for improved study quality (BRISQ).J Proteome Res. 2011 Aug 5;10(8):3429-38. doi: 10.1021/pr200021n. Epub 2011 Jun 21. J Proteome Res. 2011. PMID: 21574648 Free PMC article.
-
Biospecimen Reporting for Improved Study Quality.Biopreserv Biobank. 2011 Apr;9(1):57-70. doi: 10.1089/bio.2010.0036. Biopreserv Biobank. 2011. PMID: 21826252 Free PMC article.
-
Universal Reference RNA is not a representative normal sample for oligonucleotide microarray studies.Pathol Oncol Res. 2008 Sep;14(3):243-51. doi: 10.1007/s12253-008-9068-2. Epub 2008 Jun 14. Pathol Oncol Res. 2008. PMID: 18553159
References
-
- Emmert-Buck MR, Strausberg RL, Krizman DB, Bonaldo MF, Bonner RF, Bostwick DG, Brown MR, Buetow KH, Chuaqui RF, Cole KA, Duray PH, Englert CR, Gillespie JW, Greenhut S, Grouse L, Hillier LW, Katz KS, Klausner RD, Kuznetzov V, Lash AE, Lennon G, Linehan WM, Liotta LA, Marra MA, Munson PJ, Ornstein DK, Prabhu VV, Prange C, Schuler GD, Soares MB, Tolstoshev CM, Vocke CD, Waterston RH. Molecular profiling of clinical tissue specimens. J Mol Diagn. 2000;2:60–66. - PMC - PubMed
-
- Tureci O, Ding J, Hilton H, Bian H, Ohkawa H, Braxenthaler M, Seitz G, Raddrizzani l, Friess H, Buchler M, Sahin U, Hammer J. Computational dissection of tissue contamination for identification of colon cancer specific expression profiles. FASEB J. 2003;17:376–385. doi: 10.1096/fj.02-0478com. - DOI - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

