Human transcriptome subtraction by using short sequence tags to search for tumor viruses in conjunctival carcinoma
- PMID: 17686852
- PMCID: PMC2045575
- DOI: 10.1128/JVI.00875-07
Human transcriptome subtraction by using short sequence tags to search for tumor viruses in conjunctival carcinoma
Abstract
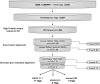
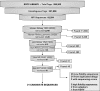
Digital transcript subtraction (DTS) was developed to subtract in silico known human sequences from expression library data sets, leaving candidate nonhuman sequences for further analysis. This approach requires precise discrimination between human and nonhuman cDNA sequences. Database comparisons show high likelihood that small viral sequences can be successfully distinguished from human sequences. DTS analysis of 9,026 20-bp tags from an expression library of BCBL-1 cells infected with Kaposi's sarcoma-associated herpesvirus (KSHV) resolved all but three candidate sequences. Two of these sequences belonged to KSHV transcripts, and the third belonged to an unannotated human expression sequence tag. Overall, 0.24% of transcripts from this cell line were of viral origin. DTS analysis of 241,122 expression tags from three squamous cell conjunctival carcinomas revealed that only 21 sequences did not align with sequences from human databases. All 21 candidates amplify human transcripts and have secondary evidence for being of human origin. This analysis shows that it is unlikely that distinguishable viral transcripts are present in conjunctival carcinomas at 20 transcripts per million or higher, which is the equivalent of approximately 4 transcripts per cell. DTS is a simple screening method to discover novel viral nucleic acids. It provides, for the first time, quantitative evidence against some classes of viral etiology when no viral transcripts are found, thereby reducing the uncertainty involved in new pathogen discovery.
Figures

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

