Identification and validation of biomarkers of IgV(H) mutation status in chronic lymphocytic leukemia using microfluidics quantitative real-time polymerase chain reaction technology
- PMID: 17690214
- PMCID: PMC1975107
- DOI: 10.2353/jmoldx.2007.070001
Identification and validation of biomarkers of IgV(H) mutation status in chronic lymphocytic leukemia using microfluidics quantitative real-time polymerase chain reaction technology
Abstract
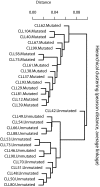
To develop a model incorporating relevant prognostic biomarkers for untreated chronic lymphocytic leukemia patients, we re-analyzed the raw data from four published gene expression profiling studies. We selected 88 candidate biomarkers linked to immunoglobulin heavy-chain variable region gene (IgV(H)) mutation status and produced a reliable and reproducible microfluidics quantitative real-time polymerase chain reaction array. We applied this array to a training set of 29 purified samples from previously untreated patients. In an unsupervised analysis, the samples clustered into two groups. Using a cutoff point of 2% homology to the germline IgV(H) sequence, one group contained all 14 IgV(H)-unmutated samples; the other contained all 15 mutated samples. We confirmed the differential expression of 37 of the candidate biomarkers using two-sample t-tests. Next, we constructed 16 different models to predict IgV(H) mutation status and evaluated their performance on an independent test set of 20 new samples. Nine models correctly classified 11 of 11 IgV(H)-mutated cases and eight of nine IgV(H)-unmutated cases, with some models using three to seven genes. Thus, we can classify cases with 95% accuracy based on the expression of as few as three genes.
Figures
Similar articles
-
ZAP-70 compared with immunoglobulin heavy-chain gene mutation status as a predictor of disease progression in chronic lymphocytic leukemia.N Engl J Med. 2004 Aug 26;351(9):893-901. doi: 10.1056/NEJMoa040857. N Engl J Med. 2004. PMID: 15329427
-
Variations in the detection of ZAP-70 in chronic lymphocytic leukemia: Comparison with IgV(H) mutation analysis.Cytometry B Clin Cytom. 2006 Jul 15;70(4):270-5. doi: 10.1002/cyto.b.20134. Cytometry B Clin Cytom. 2006. PMID: 16906585
-
Biological validation of differentially expressed genes in chronic lymphocytic leukemia identified by applying multiple statistical methods to oligonucleotide microarrays.J Mol Diagn. 2005 Aug;7(3):337-45. doi: 10.1016/s1525-1578(10)60562-4. J Mol Diagn. 2005. PMID: 16049305 Free PMC article.
-
Molecular pathogenesis of chronic lymphocytic leukemia.J Clin Invest. 2012 Oct;122(10):3432-8. doi: 10.1172/JCI64101. Epub 2012 Oct 1. J Clin Invest. 2012. PMID: 23023714 Free PMC article. Review.
-
Incorporating prognostic information into treatment decisions in chronic lymphocytic leukemia.Curr Oncol Rep. 2009 Sep;11(5):353-9. doi: 10.1007/s11912-009-0048-9. Curr Oncol Rep. 2009. PMID: 19679010 Review.
Cited by
-
Cross-species transcriptional network analysis defines shared inflammatory responses in murine and human lupus nephritis.J Immunol. 2012 Jul 15;189(2):988-1001. doi: 10.4049/jimmunol.1103031. Epub 2012 Jun 20. J Immunol. 2012. PMID: 22723521 Free PMC article.
-
Cryptochrome-1 Gene Expression is a Reliable Prognostic Indicator in Egyptian Patients with Chronic Lymphocytic Leukemia: A Prospective Cohort Study.Turk J Haematol. 2018 Aug 3;35(3):168-174. doi: 10.4274/tjh.2017.0169. Epub 2017 Sep 8. Turk J Haematol. 2018. PMID: 28884705 Free PMC article.
-
Regulatory network reconstruction reveals genes with prognostic value for chronic lymphocytic leukemia.BMC Genomics. 2015 Nov 25;16:1002. doi: 10.1186/s12864-015-2189-6. BMC Genomics. 2015. PMID: 26606983 Free PMC article.
-
A gene expression assay based on chronic lymphocytic leukemia activation in the microenvironment to predict progression.Blood Adv. 2022 Nov 8;6(21):5763-5773. doi: 10.1182/bloodadvances.2022007508. Blood Adv. 2022. PMID: 35973197 Free PMC article.
-
LDOC1 mRNA is differentially expressed in chronic lymphocytic leukemia and predicts overall survival in untreated patients.Blood. 2011 Apr 14;117(15):4076-84. doi: 10.1182/blood-2010-09-304881. Epub 2011 Feb 10. Blood. 2011. PMID: 21310924 Free PMC article.
References
-
- Rai K, Patel D. Chronic lymphocytic leukemia. Hematology: Basic Principles and Practice. Hoffman R, Benz E Jr, Shattil S, Furie B, Cohen H, Silberstein L, McGlave P, editors. New York: Churchill Livingstone; 2000
-
- Hallek M. New concepts in the pathogenesis, diagnosis, prognostic factors, and clinical presentation of chronic lymphocytic leukemia. Rev Clin Exp Hematol. 2000;4:103–117.
-
- Keating MJ, O’Brien S. Conventional management of chronic lymphocytic leukemia. Rev Clin Exp Hematol. 2000;4:118–133. - PubMed
-
- Hamblin TJ, Davis Z, Gardiner A, Oscier DG, Stevenson FK. Unmutated Ig V(H) genes are associated with a more aggressive form of chronic lymphocytic leukemia. Blood. 1999;94:1848–1854. - PubMed
-
- Damle RN, Wasil T, Fais F, Ghiotto F, Valetto A, Allen SL, Buchbinder A, Budman D, Dittmar K, Kolitz J, Lichtman SM, Schulman P, Vinciguerra VP, Rai KR, Ferrarini M, Chiorazzi N. Ig V gene mutation status and CD38 expression as novel prognostic indicators in chronic lymphocytic leukemia. Blood. 1999;94:1840–1847. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources