Characterization and biological role of the O-polysaccharide gene cluster of Yersinia enterocolitica serotype O:9
- PMID: 17693522
- PMCID: PMC2168460
- DOI: 10.1128/JB.00605-07
Characterization and biological role of the O-polysaccharide gene cluster of Yersinia enterocolitica serotype O:9
Abstract
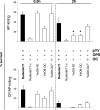
Yersinia enterocolitica serotype O:9 is a gram-negative enteropathogen that infects animals and humans. The role of lipopolysaccharide (LPS) in Y. enterocolitica O:9 pathogenesis, however, remains unclear. The O:9 LPS consists of lipid A to which is linked the inner core oligosaccharide, serving as an attachment site for both the outer core (OC) hexasaccharide and the O-polysaccharide (OPS; a homopolymer of N-formylperosamine). In this work, we cloned the OPS gene cluster of O:9 and identified 12 genes organized into four operons upstream of the gnd gene. Ten genes were predicted to encode glycosyltransferases, the ATP-binding cassette polysaccharide translocators, or enzymes required for the biosynthesis of GDP-N-formylperosamine. The two remaining genes within the OPS gene cluster, galF and galU, were not ascribed a clear function in OPS biosynthesis; however, the latter gene appeared to be essential for O:9. The biological functions of O:9 OPS and OC were studied using isogenic mutants lacking one or both of these LPS parts. We showed that OPS and OC confer resistance to human complement and polymyxin B; the OPS effect on polymyxin B resistance could be observed only in the absence of OC.
Figures


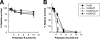

Similar articles
-
Characterization of the six glycosyltransferases involved in the biosynthesis of Yersinia enterocolitica serotype O:3 lipopolysaccharide outer core.J Biol Chem. 2010 Sep 3;285(36):28333-42. doi: 10.1074/jbc.M110.111336. Epub 2010 Jul 1. J Biol Chem. 2010. PMID: 20595390 Free PMC article.
-
A novel locus of Yersinia enterocolitica serotype O:3 involved in lipopolysaccharide outer core biosynthesis.Mol Microbiol. 1995 Aug;17(3):575-94. doi: 10.1111/j.1365-2958.1995.mmi_17030575.x. Mol Microbiol. 1995. PMID: 8559076
-
Identification of three oligo-/polysaccharide-specific ligases in Yersinia enterocolitica.Mol Microbiol. 2012 Jan;83(1):125-36. doi: 10.1111/j.1365-2958.2011.07918.x. Epub 2011 Nov 28. Mol Microbiol. 2012. PMID: 22053911
-
Yersinia enterocolitica lipopolysaccharide: genetics and virulence.Trends Microbiol. 1993 Jul;1(4):148-52. doi: 10.1016/0966-842x(93)90130-j. Trends Microbiol. 1993. PMID: 7511477 Review.
-
The biosynthesis and biological role of lipopolysaccharide O-antigens of pathogenic Yersiniae.Carbohydr Res. 2003 Nov 14;338(23):2521-9. doi: 10.1016/s0008-6215(03)00305-7. Carbohydr Res. 2003. PMID: 14670713 Review.
Cited by
-
Exploiting the Campylobacter jejuni protein glycosylation system for glycoengineering vaccines and diagnostic tools directed against brucellosis.Microb Cell Fact. 2012 Jan 25;11:13. doi: 10.1186/1475-2859-11-13. Microb Cell Fact. 2012. PMID: 22276812 Free PMC article.
-
Regulatory protein OmpR influences the serum resistance of Yersinia enterocolitica O:9 by modifying the structure of the outer membrane.PLoS One. 2013 Nov 19;8(11):e79525. doi: 10.1371/journal.pone.0079525. eCollection 2013. PLoS One. 2013. PMID: 24260242 Free PMC article.
-
A transposon site hybridization screen identifies galU and wecBC as important for survival of Yersinia pestis in murine macrophages.J Bacteriol. 2012 Feb;194(3):653-62. doi: 10.1128/JB.06237-11. Epub 2011 Dec 2. J Bacteriol. 2012. PMID: 22139502 Free PMC article.
-
Genetics and evolution of Yersinia pseudotuberculosis O-specific polysaccharides: a novel pattern of O-antigen diversity.FEMS Microbiol Rev. 2017 Mar 1;41(2):200-217. doi: 10.1093/femsre/fux002. FEMS Microbiol Rev. 2017. PMID: 28364730 Free PMC article. Review.
-
A bioconjugate vaccine against Brucella abortus produced by engineered Escherichia coli.Front Bioeng Biotechnol. 2023 Feb 23;11:1121074. doi: 10.3389/fbioe.2023.1121074. eCollection 2023. Front Bioeng Biotechnol. 2023. PMID: 36911199 Free PMC article.
References
-
- Al-Hendy, A., P. Toivanen, and M. Skurnik. 1991. The effect of growth temperature on the biosynthesis of Yersinia enterocolitica O:3 lipopolysaccharide: temperature regulates the transcription of the rfb but not of the rfa region. Microb. Pathog. 10:81-86. - PubMed
-
- Al-Hendy, A., P. Toivanen, and M. Skurnik. 1991. Expression cloning of Yersinia enterocolitica O:3 rfb gene cluster in Escherichia coli K12. Microb. Pathog. 10:47-59. - PubMed
-
- Altschul, S. F., W. Gish, W. Miller, E. W. Myers, and D. J. Lipman. 1990. Basic local alignment search tool. J. Mol. Biol. 215:403-410. - PubMed
-
- Anisimov, A. P., S. V. Dentovskaya, G. M. Titareva, I. V. Bakhteeva, R. Z. Shaikhutdinova, S. V. Balakhonov, B. Lindner, N. A. Kocharova, S. N. Senchenkova, O. Holst, G. B. Pier, and Y. A. Knirel. 2005. Intraspecies and temperature-dependent variations in susceptibility of Yersinia pestis to the bactericidal action of serum and to polymyxin B. Infect. Immun. 73:7324-7331. - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources

