The maize floury1 gene encodes a novel endoplasmic reticulum protein involved in zein protein body formation
- PMID: 17693529
- PMCID: PMC2002605
- DOI: 10.1105/tpc.107.053538
The maize floury1 gene encodes a novel endoplasmic reticulum protein involved in zein protein body formation
Abstract
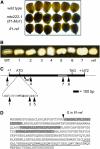
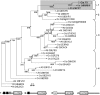
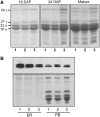
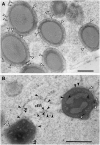
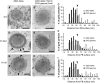
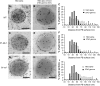
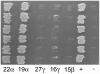
The maize (Zea mays) floury1 (fl1) mutant was first reported almost 100 years ago, but its molecular identity has remained unknown. We report the cloning of Fl1, which encodes a novel zein protein body membrane protein with three predicted transmembrane domains and a C-terminal plant-specific domain of unknown function (DUF593). In wild-type endosperm, the FL1 protein accumulates at a high level during the period of zein synthesis and protein body development and declines to a low level at kernel maturity. Immunogold labeling showed that FL1 resides in the endoplasmic reticulum surrounding the protein body. Zein protein bodies in fl1 mutants are of normal size, shape, and abundance. However, mutant protein bodies ectopically accumulate 22-kD alpha-zeins in the gamma-zein-rich periphery and center of the core, rather than their normal discrete location in a ring at outer edge of the core. The 19-kD alpha-zein is uniformly distributed throughout the core in wild-type protein bodies, and this distribution is unaffected in fl1 mutants. Pairwise yeast two-hybrid experiments showed that FL1 DUF593 interacts with the 22-kD alpha-zein. Results of these studies suggest that FL1 participates in protein body formation by facilitating the localization of 22-kD alpha-zein and that this is essential for the formation of vitreous endosperm.
Figures



Similar articles
-
Zein protein interactions, rather than the asymmetric distribution of zein mRNAs on endoplasmic reticulum membranes, influence protein body formation in maize endosperm.Plant Cell. 2002 Mar;14(3):655-72. doi: 10.1105/tpc.010431. Plant Cell. 2002. PMID: 11910012 Free PMC article.
-
Asymmetric localization of seed storage protein RNAs to distinct subdomains of the endoplasmic reticulum in developing maize endosperm cells.Plant Cell Physiol. 2004 Dec;45(12):1830-7. doi: 10.1093/pcp/pch210. Plant Cell Physiol. 2004. PMID: 15653801
-
Opaque1 encodes a myosin XI motor protein that is required for endoplasmic reticulum motility and protein body formation in maize endosperm.Plant Cell. 2012 Aug;24(8):3447-62. doi: 10.1105/tpc.112.101360. Epub 2012 Aug 14. Plant Cell. 2012. PMID: 22892319 Free PMC article.
-
The regulation of zein biosynthesis in maize endosperm.Theor Appl Genet. 2020 May;133(5):1443-1453. doi: 10.1007/s00122-019-03520-z. Epub 2020 Jan 2. Theor Appl Genet. 2020. PMID: 31897513 Review.
-
Molecular mechanisms regulating the synthesis of storage proteins in maize endosperm.Crit Rev Food Sci Nutr. 1982;16(2):199-215. doi: 10.1080/10408398209527334. Crit Rev Food Sci Nutr. 1982. PMID: 7039978 Review. No abstract available.
Cited by
-
Internal membranes in maize aleurone protein storage vacuoles: beyond autophagy.Plant Cell. 2011 Dec;23(12):4168-71; author reply 4171-2. doi: 10.1105/tpc.111.092551. Epub 2011 Dec 16. Plant Cell. 2011. PMID: 22180629 Free PMC article. No abstract available.
-
Comparative transcriptomics reveals the difference in early endosperm development between maize with different amylose contents.PeerJ. 2019 Aug 28;7:e7528. doi: 10.7717/peerj.7528. eCollection 2019. PeerJ. 2019. PMID: 31523504 Free PMC article.
-
Specialized endoplasmic reticulum-derived vesicles in plants: Functional diversity, evolution, and biotechnological exploitation.J Integr Plant Biol. 2022 Apr;64(4):821-835. doi: 10.1111/jipb.13233. Epub 2022 Mar 9. J Integr Plant Biol. 2022. PMID: 35142108 Free PMC article. Review.
-
Kinesins and Myosins: Molecular Motors that Coordinate Cellular Functions in Plants.Annu Rev Plant Biol. 2018 Apr 29;69:329-361. doi: 10.1146/annurev-arplant-042817-040024. Epub 2018 Feb 28. Annu Rev Plant Biol. 2018. PMID: 29489391 Free PMC article. Review.
-
Protein body formation in the endoplasmic reticulum as an evolution of storage protein sorting to vacuoles: insights from maize γ-zein.Front Plant Sci. 2014 Jul 15;5:331. doi: 10.3389/fpls.2014.00331. eCollection 2014. Front Plant Sci. 2014. PMID: 25076952 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

