Genotypic evolution of azole resistance mechanisms in sequential Candida albicans isolates
- PMID: 17693596
- PMCID: PMC2043391
- DOI: 10.1128/EC.00151-07
Genotypic evolution of azole resistance mechanisms in sequential Candida albicans isolates
Abstract
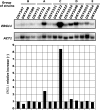
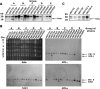
TAC1 (for transcriptional activator of CDR genes) is critical for the upregulation of the ABC transporters CDR1 and CDR2, which mediate azole resistance in Candida albicans. While a wild-type TAC1 allele drives high expression of CDR1/2 in response to inducers, we showed previously that TAC1 can be hyperactive by a gain-of-function (GOF) point mutation responsible for constitutive high expression of CDR1/2. High azole resistance levels are achieved when C. albicans carries hyperactive alleles only as a consequence of loss of heterozygosity (LOH) at the TAC1 locus on chromosome 5 (Chr 5), which is linked to the mating-type-like (MTL) locus. Both are located on the Chr 5 left arm along with ERG11 (target of azoles). In this work, five groups of related isolates containing azole-susceptible and -resistant strains were analyzed for the TAC1 and ERG11 alleles and for Chr 5 alterations. While recovered ERG11 alleles contained known mutations, 17 new TAC1 alleles were isolated, including 7 hyperactive alleles with five separate new GOF mutations. Single-nucleotide-polymorphism analysis of Chr 5 revealed that azole-resistant strains acquired TAC1 hyperactive alleles and, in most cases, ERG11 mutant alleles by LOH events not systematically including the MTL locus. TAC1 LOH resulted from mitotic recombination of the left arm of Chr 5, gene conversion within the TAC1 locus, or the loss and reduplication of the entire Chr 5. In one case, two independent TAC1 hyperactive alleles were acquired. Comparative genome hybridization and karyotype analysis revealed the presence of isochromosome 5L [i(5L)] in two azole-resistant strains. i(5L) leads to increased copy numbers of azole resistance genes present on the left arm of Chr 5, among them TAC1 and ERG11. Our work shows that azole resistance was due not only to the presence of specific mutations in azole resistance genes (at least ERG11 and TAC1) but also to their increase in copy number by LOH and to the addition of extra Chr 5 copies. With the combination of these different modifications, sophisticated genotypes were obtained. The development of azole resistance in C. albicans is therefore a powerful instrument for generating genetic diversity.
Figures






References
-
- Akins, R. A. 2005. An update on antifungal targets and mechanisms of resistance in Candida albicans. Med. Mycol. 43:285-318. - PubMed
-
- Andaluz, E., T. Ciudad, J. Gomez-Raja, R. Calderone, and G. Larriba. 2006. Rad52 depletion in Candida albicans triggers both the DNA-damage checkpoint and filamentation accompanied by but independent of expression of hypha-specific genes. Mol. Microbiol. 59:1452-1472. - PubMed
-
- Barker, K. S., and P. D. Rogers. 2006. Recent insights into the mechanisms of antifungal resistance. Curr. Infect. Dis. Rep. 8:449-456. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

